Asynchronous combinatorial action of four regulatory factors activates Bcl11b for T cell commitment
- PMID: 27376470
- PMCID: PMC4955789
- DOI: 10.1038/ni.3514
Asynchronous combinatorial action of four regulatory factors activates Bcl11b for T cell commitment
Abstract
During T cell development, multipotent progenitors relinquish competence for other fates and commit to the T cell lineage by turning on Bcl11b, which encodes a transcription factor. To clarify lineage commitment mechanisms, we followed developing T cells at the single-cell level using Bcl11b knock-in fluorescent reporter mice. Notch signaling and Notch-activated transcription factors collaborate to activate Bcl11b expression irrespectively of Notch-dependent proliferation. These inputs work via three distinct, asynchronous mechanisms: an early locus 'poising' function dependent on TCF-1 and GATA-3, a stochastic-permissivity function dependent on Notch signaling, and a separate amplitude-control function dependent on Runx1, a factor already present in multipotent progenitors. Despite their necessity for Bcl11b expression, these inputs act in a stage-specific manner, providing a multitiered mechanism for developmental gene regulation.
Figures




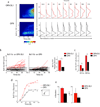



Comment in
-
Induction of Bcl11b during T cell commitment through a tripartite mechanism.Nat Immunol. 2016 Jul 19;17(8):903-4. doi: 10.1038/ni.3520. Nat Immunol. 2016. PMID: 27434004 No abstract available.
References
Reference List
-
- Liu P, Li P, Burke S. Critical roles of Bcl11b in T-cell development and maintenance of T-cell identity. Immunol Rev. 2010;238:138–149. - PubMed
METHODS-ONLY REFERENCES
-
- Hernandez-Hoyos G, Anderson MK, Wang C, Rothenberg EV, Alberola-Ila J. GATA-3 expression is controlled by TCR signals and regulates CD4/CD8 differentiation. Immunity. 2003;19:83–94. - PubMed
-
- Telfer JC, Hedblom EE, Anderson MK, Laurent MN, Rothenberg EV. Localization of the domains in Runx transcription factors required for the repression of CD4 in thymocytes. J Immunol. 2004;172:4359–4370. - PubMed
-
- Nakano T, Kodama H, Honjo T. Generation of lymphohematopoietic cells from embryonic stem cells in culture. Science. 1994;265:1098–1101. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

