Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment
- PMID: 27378076
- PMCID: PMC4935971
- DOI: 10.1038/ncomms12115
Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment
Abstract
Biological features can be inferred, based on genomic data, for many microbial lineages that remain uncultured. However, cultivation is important for characterizing an organism's physiology and testing its genome-encoded potential. Here we use single-cell genomics to infer cultivation conditions for the isolation of an ectosymbiotic Nanoarchaeota ('Nanopusillus acidilobi') and its host (Acidilobus, a crenarchaeote) from a terrestrial geothermal environment. The cells of 'Nanopusillus' are among the smallest known cellular organisms (100-300 nm). They appear to have a complete genetic information processing machinery, but lack almost all primary biosynthetic functions as well as respiration and ATP synthesis. Genomic and proteomic comparison with its distant relative, the marine Nanoarchaeum equitans illustrate an ancient, common evolutionary history of adaptation of the Nanoarchaeota to ectosymbiosis, so far unique among the Archaea.
Figures





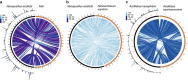

References
-
- Brown C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015). - PubMed
-
- Rinke C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013). - PubMed
-
- Wrighton K. C. et al. Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337, 1661–1665 (2012). - PubMed
-
- Castelle C. J. et al. Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr. Biol. 25, 690–701 (2015). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

