MSK1 triggers the expression of the INK4AB/ARF locus in oncogene-induced senescence
- PMID: 27385346
- PMCID: PMC5007092
- DOI: 10.1091/mbc.E15-11-0772
MSK1 triggers the expression of the INK4AB/ARF locus in oncogene-induced senescence
Abstract
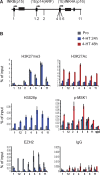
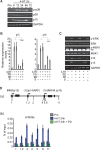
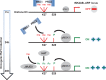
The tumor suppressor proteins p15(INK4B), p16(INK4A), and p14(ARF), encoded by the INK4AB/ARF locus, are crucial regulators of cellular senescence. The locus is epigenetically silenced by the repressive Polycomb complexes in growing cells but is activated in response to oncogenic stress. Here we show that the mitogen- and stress-activated kinase (MSK1) is up-regulated after RAF1 oncogenic stress and that the phosphorylated (activated) form of MSK1 is significantly increased in the nucleus and recruited to the INK4AB/ARF locus. We show that MSK1 mediates histone H3S28 phosphorylation at the INK4AB/ARF locus and contributes to the rapid transcriptional activation of p15(INK4B) and p16(INK4A) in human cells despite the presence of the repressive H3K27me3 mark. Furthermore, we show that upon MSK1 depletion in oncogenic RAF1-expressing cells, H3S28ph presence at the INK4 locus and p15(INK4B) and p16(INK4A) expression are reduced. Finally, we show that H3S28-MSK-dependent phosphorylation functions in response to RAF1 signaling and that ERK and p38α contribute to MSK1 activation in oncogene-induced senescence.
© 2016 Culerrier et al. This article is distributed by The American Society for Cell Biology under license from the author(s). Two months after publication it is available to the public under an Attribution–Noncommercial–Share Alike 3.0 Unported Creative Commons License (http://creativecommons.org/licenses/by-nc-sa/3.0).
Figures



Similar articles
-
Enhancer of zeste homolog 2 depletion induces cellular senescence via histone demethylation along the INK4/ARF locus.Int J Biochem Cell Biol. 2015 Aug;65:104-12. doi: 10.1016/j.biocel.2015.05.011. Epub 2015 May 22. Int J Biochem Cell Biol. 2015. PMID: 26004298
-
Prognostic significance of p14ARF, p15INK4b, and p16INK4a inactivation in malignant peripheral nerve sheath tumors.Clin Cancer Res. 2011 Jun 1;17(11):3771-82. doi: 10.1158/1078-0432.CCR-10-2393. Epub 2011 Jan 24. Clin Cancer Res. 2011. PMID: 21262917
-
Polycomb mediated epigenetic silencing and replication timing at the INK4a/ARF locus during senescence.PLoS One. 2009 May 20;4(5):e5622. doi: 10.1371/journal.pone.0005622. PLoS One. 2009. PMID: 19462008 Free PMC article.
-
Long noncoding RNA, polycomb, and the ghosts haunting INK4b-ARF-INK4a expression.Cancer Res. 2011 Aug 15;71(16):5365-9. doi: 10.1158/0008-5472.CAN-10-4379. Epub 2011 Aug 9. Cancer Res. 2011. PMID: 21828241 Free PMC article. Review.
-
Deletion of RDINK4/ARF enhancer: A novel mutation to "inactivate" the INK4-ARF locus.DNA Repair (Amst). 2017 Sep;57:50-55. doi: 10.1016/j.dnarep.2017.06.027. Epub 2017 Jun 24. DNA Repair (Amst). 2017. PMID: 28688373 Review.
Cited by
-
Circular ANRIL isoforms switch from repressors to activators of p15/CDKN2B expression during RAF1 oncogene-induced senescence.RNA Biol. 2021 Mar;18(3):404-420. doi: 10.1080/15476286.2020.1812910. Epub 2020 Sep 29. RNA Biol. 2021. PMID: 32862732 Free PMC article.
-
Regulation of senescence traits by MAPKs.Geroscience. 2020 Apr;42(2):397-408. doi: 10.1007/s11357-020-00183-3. Epub 2020 Apr 16. Geroscience. 2020. PMID: 32300964 Free PMC article. Review.
References
-
- Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, Jones RS, Zhang Y. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science. 2002;298:1039–1043. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

