Strategies for Comparing Metabolic Profiles: Implications for the Inference of Biochemical Mechanisms from Metabolomics Data
- PMID: 27392364
- PMCID: PMC5708160
- DOI: 10.1109/TCBB.2016.2586065
Strategies for Comparing Metabolic Profiles: Implications for the Inference of Biochemical Mechanisms from Metabolomics Data
Abstract
Background: Large amounts of metabolomics data have been accumulated in recent years and await analysis. Previously, we had developed a systems biology approach to infer biochemical mechanisms underlying metabolic alterations observed in cancers and other diseases. The method utilized the typical Euclidean distance for comparing metabolic profiles. Here, we ask whether any of the numerous alternative metrics might serve this purpose better.
Methods and findings: We used enzymatic alterations in purine metabolism that were measured in human renal cell carcinoma to test various metrics with the goal of identifying the best metrics for discerning metabolic profiles of healthy and diseased individuals. The results showed that several metrics have similarly good performance, but that some are unsuited for comparisons of metabolic profiles. Furthermore, the results suggest that relative changes in metabolite levels, which reduce bias toward large metabolite concentrations, are better suited for comparisons of metabolic profiles than absolute changes. Finally, we demonstrate that a sequential search for enzymatic alterations, ranked by importance, is not always valid.
Conclusions: We identified metrics that are appropriate for comparisons of metabolic profiles. In addition, we constructed strategic guidelines for the algorithmic identification of biochemical mechanisms from metabolomics data.
Conflict of interest statement
The authors have no conflict of interest to declare.
Figures
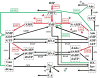
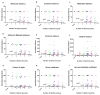


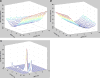


Similar articles
-
Inference of cancer mechanisms through computational systems analysis.Mol Biosyst. 2017 Feb 28;13(3):489-497. doi: 10.1039/c6mb00672h. Mol Biosyst. 2017. PMID: 28112324 Free PMC article.
-
Unveiling cellular biochemical reactions via metabolomics-driven approaches.Curr Opin Microbiol. 2010 Jun;13(3):358-62. doi: 10.1016/j.mib.2010.04.006. Epub 2010 Apr 27. Curr Opin Microbiol. 2010. PMID: 20430690 Review.
-
Simulation and Reconstruction of Metabolite-Metabolite Association Networks Using a Metabolic Dynamic Model and Correlation Based Algorithms.J Proteome Res. 2019 Mar 1;18(3):1099-1113. doi: 10.1021/acs.jproteome.8b00781. Epub 2019 Feb 4. J Proteome Res. 2019. PMID: 30663881
-
Informatics for Metabolomics.Adv Exp Med Biol. 2016;939:91-115. doi: 10.1007/978-981-10-1503-8_5. Adv Exp Med Biol. 2016. PMID: 27807745
-
Metabolome 2.0: quantitative genetics and network biology of metabolic phenotypes.Mol Biosyst. 2012 Oct;8(10):2494-502. doi: 10.1039/c2mb25167a. Mol Biosyst. 2012. PMID: 22868675 Review.
Cited by
-
Delineating the Role of the Urinary Metabolome in the Lithogenesis of Calcium-Based Kidney Stones.Urology. 2022 Sep;167:49-55. doi: 10.1016/j.urology.2022.06.004. Epub 2022 Jun 15. Urology. 2022. PMID: 35716870 Free PMC article.
-
Complex system modeling reveals oxalate homeostasis is driven by diverse oxalate-degrading bacteria.Elife. 2025 May 1;14:RP104121. doi: 10.7554/eLife.104121. Elife. 2025. PMID: 40310467 Free PMC article.
-
Learning a confidence score and the latent space of a new supervised autoencoder for diagnosis and prognosis in clinical metabolomic studies.BMC Bioinformatics. 2022 Sep 1;23(1):361. doi: 10.1186/s12859-022-04900-x. BMC Bioinformatics. 2022. PMID: 36050631 Free PMC article.
-
Targeted metabolomics characterizes metabolite occurrence and variability in stable freshwater mussel populations.Conserv Physiol. 2023 Jun 10;11(1):coad040. doi: 10.1093/conphys/coad040. eCollection 2023. Conserv Physiol. 2023. PMID: 37701372 Free PMC article.
-
Phenotypic and metabolomic characteristics of mouse models of metabolic associated steatohepatitis.Biomark Res. 2024 Jan 9;12(1):6. doi: 10.1186/s40364-023-00555-9. Biomark Res. 2024. PMID: 38195587 Free PMC article.
References
-
- Wu W, Zhao S. Metabolic changes in cancer: beyond the Warburg effect. Acta Biochim Biophys Sin (Shanghai) 2013 Jan;45(1):18–26. - PubMed
-
- Schulze A, Harris AL. How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature. 2012 Nov 15;491(7424):364–73. - PubMed
-
- Park FC. Distance Metrics on the Rigid-Body Motions with Applications to Mechanism Design. Journal of Mechanical Design. 1995;117(1):48–54.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

