An ancestral stomatal patterning module revealed in the non-vascular land plant Physcomitrella patens
- PMID: 27407102
- PMCID: PMC5047656
- DOI: 10.1242/dev.135038
An ancestral stomatal patterning module revealed in the non-vascular land plant Physcomitrella patens
Abstract
The patterning of stomata plays a vital role in plant development and has emerged as a paradigm for the role of peptide signals in the spatial control of cellular differentiation. Research in Arabidopsis has identified a series of epidermal patterning factors (EPFs), which interact with an array of membrane-localised receptors and associated proteins (encoded by ERECTA and TMM genes) to control stomatal density and distribution. However, although it is well-established that stomata arose very early in the evolution of land plants, until now it has been unclear whether the established angiosperm stomatal patterning system represented by the EPF/TMM/ERECTA module reflects a conserved, universal mechanism in the plant kingdom. Here, we use molecular genetics to show that the moss Physcomitrella patens has conserved homologues of angiosperm EPF, TMM and at least one ERECTA gene that function together to permit the correct patterning of stomata and that, moreover, elements of the module retain function when transferred to Arabidopsis Our data characterise the stomatal patterning system in an evolutionarily distinct branch of plants and support the hypothesis that the EPF/TMM/ERECTA module represents an ancient patterning system.
Keywords: Evolution; Patterning; Peptide signalling; Stomata.
© 2016. Published by The Company of Biologists Ltd.
Conflict of interest statement
The authors declare no competing or financial interests.
Figures

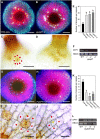
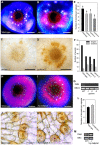
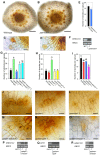
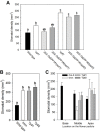
References
-
- Beerling D. J. (2007). The Emerald Planet: How Plants Changed Earth's History. Oxford: Oxford University Press.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

