Peptidyl-Proline Isomerases (PPIases): Targets for Natural Products and Natural Product-Inspired Compounds
- PMID: 27409354
- PMCID: PMC5501181
- DOI: 10.1021/acs.jmedchem.6b00411
Peptidyl-Proline Isomerases (PPIases): Targets for Natural Products and Natural Product-Inspired Compounds
Abstract
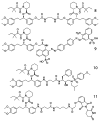
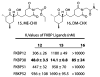
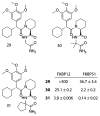
Peptidyl-proline isomerases (PPIases) are a chaperone superfamily comprising the FK506-binding proteins (FKBPs), cyclophilins, and parvulins. PPIases catalyze the cis/trans isomerization of proline, acting as a regulatory switch during folding, activation, and/or degradation of many proteins. These "clients" include proteins with key roles in cancer, neurodegeneration, and psychiatric disorders, suggesting that PPIase inhibitors could be important therapeutics. However, the active site of PPIases is shallow, solvent-exposed, and well conserved between family members, making selective inhibitor design challenging. Despite these hurdles, macrocyclic natural products, including FK506, rapamycin, and cyclosporin, bind PPIases with nanomolar or better affinity. De novo attempts to derive new classes of inhibitors have been somewhat less successful, often showcasing the "undruggable" features of PPIases. Interestingly, the most potent of these next-generation molecules tend to integrate features of the natural products, including macrocyclization or proline mimicry strategies. Here, we review recent developments and ongoing challenges in the inhibition of PPIases, with a focus on how natural products might inform the creation of potent and selective inhibitors.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
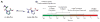

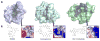

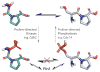
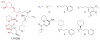

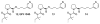


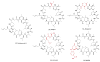
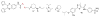
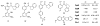

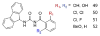
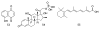
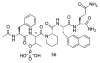
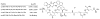
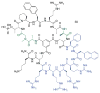

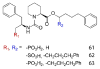
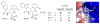

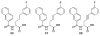
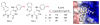

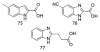
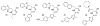
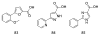
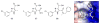
Similar articles
-
Peptidyl-prolyl cis-trans isomerases, a superfamily of ubiquitous folding catalysts.Cell Mol Life Sci. 1999 Mar;55(3):423-36. doi: 10.1007/s000180050299. Cell Mol Life Sci. 1999. PMID: 10228556 Free PMC article. Review.
-
Structural insights into Plasmodium PPIases.Front Cell Infect Microbiol. 2022 Sep 2;12:931635. doi: 10.3389/fcimb.2022.931635. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36118020 Free PMC article. Review.
-
Pharmacological targeting of catalyzed protein folding: the example of peptide bond cis/trans isomerases.Handb Exp Pharmacol. 2006;(172):359-404. doi: 10.1007/3-540-29717-0_15. Handb Exp Pharmacol. 2006. PMID: 16610367 Review.
-
Biosynthesis of cyclosporins and other natural peptidyl prolyl cis/trans isomerase inhibitors.Biochim Biophys Acta. 2015 Oct;1850(10):2111-20. doi: 10.1016/j.bbagen.2014.12.009. Epub 2014 Dec 11. Biochim Biophys Acta. 2015. PMID: 25497210 Review.
-
Peptidylprolyl cis/trans isomerases (immunophilins): biological diversity--targets--functions.Curr Top Med Chem. 2003;3(12):1315-47. doi: 10.2174/1568026033451862. Curr Top Med Chem. 2003. PMID: 12871165 Review.
Cited by
-
A novel proteotoxic combination therapy for EGFR+ and HER2+ cancers.Oncogene. 2019 May;38(22):4264-4282. doi: 10.1038/s41388-019-0717-6. Epub 2019 Feb 4. Oncogene. 2019. PMID: 30718919
-
Calcium-binding protein CALU-1 is essential for proper collagen formation in Caenorhabditis elegans.Cell Mol Life Sci. 2025 Jan 25;82(1):62. doi: 10.1007/s00018-025-05582-3. Cell Mol Life Sci. 2025. PMID: 39862239 Free PMC article.
-
Structure and function of the TPR-domain immunophilins FKBP51 and FKBP52 in normal physiology and disease.J Cell Biochem. 2024 Dec;125(12):e30406. doi: 10.1002/jcb.30406. Epub 2023 Apr 23. J Cell Biochem. 2024. PMID: 37087733 Review.
-
A Pan-Cyclophilin Inhibitor, CRV431, Decreases Fibrosis and Tumor Development in Chronic Liver Disease Models.J Pharmacol Exp Ther. 2019 Nov;371(2):231-241. doi: 10.1124/jpet.119.261099. Epub 2019 Aug 12. J Pharmacol Exp Ther. 2019. PMID: 31406003 Free PMC article.
-
Macrocyclic FKBP51 Ligands Define a Transient Binding Mode with Enhanced Selectivity.Angew Chem Int Ed Engl. 2021 Jun 7;60(24):13257-13263. doi: 10.1002/anie.202017352. Epub 2021 May 7. Angew Chem Int Ed Engl. 2021. PMID: 33843131 Free PMC article.
References
-
- Fischer G, Bang H, Mech C. Determination of enzymatic catalysis for the cis-trans-isomerization of peptide binding in proline-containing peptides. Biomed Biochim Acta. 1984;43:1101–1111. - PubMed
-
- Galat A. Peptidylprolyl cis/trans isomerases (immunophilins): Biological diversity–targets–functions. Curr Top Med Chem. 2003;3:1315–1347. - PubMed
-
- Theuerkorn M, Fischer G, Schiene-Fischer C. Prolyl cis/trans isomerase signalling pathways in cancer. Curr Opin Pharmacol. 2011;11:281–287. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information
Molecular Biology Databases

