Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation
- PMID: 27409812
- PMCID: PMC4961583
- DOI: 10.1038/nature18636
Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation
Abstract
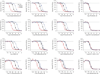
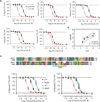
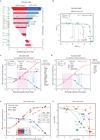
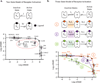
G-protein-coupled receptors (GPCRs) modulate many physiological processes by transducing a variety of extracellular cues into intracellular responses. Ligand binding to an extracellular orthosteric pocket propagates conformational change to the receptor cytosolic region to promote binding and activation of downstream signalling effectors such as G proteins and β-arrestins. It is well known that different agonists can share the same binding pocket but evoke unique receptor conformations leading to a wide range of downstream responses (‘efficacy’). Furthermore, increasing biophysical evidence, primarily using the β2-adrenergic receptor (β2AR) as a model system, supports the existence of multiple active and inactive conformational states. However, how agonists with varying efficacy modulate these receptor states to initiate cellular responses is not well understood. Here we report stabilization of two distinct β2AR conformations using single domain camelid antibodies (nanobodies)—a previously described positive allosteric nanobody (Nb80) and a newly identified negative allosteric nanobody (Nb60). We show that Nb60 stabilizes a previously unappreciated low-affinity receptor state which corresponds to one of two inactive receptor conformations as delineated by X-ray crystallography and NMR spectroscopy. We find that the agonist isoprenaline has a 15,000-fold higher affinity for β2AR in the presence of Nb80 compared to the affinity of isoprenaline for β2AR in the presence of Nb60, highlighting the full allosteric range of a GPCR. Assessing the binding of 17 ligands of varying efficacy to the β2AR in the absence and presence of Nb60 or Nb80 reveals large ligand-specific effects that can only be explained using an allosteric model which assumes equilibrium amongst at least three receptor states. Agonists generally exert efficacy by stabilizing the active Nb80-stabilized receptor state (R80). In contrast, for a number of partial agonists, both stabilization of R80 and destabilization of the inactive, Nb60-bound state (R60) contribute to their ability to modulate receptor activation. These data demonstrate that ligands can initiate a wide range of cellular responses by differentially stabilizing multiple receptor states.
Figures








References
References for Methods section
-
- Kobilka BK. Amino and carboxyl terminal modifications to facilitate the production and purification of a G protein-coupled receptor. Analytical biochemistry. 1995;231:269–271. - PubMed
-
- Otwinowski ZMW. In: Methods in Enzymology. Carter Charlles W, editor. Vol. 276. Academic Press; 1997. pp. 307–326. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

