Sleep Deprivation Influences Circadian Gene Expression in the Lateral Habenula
- PMID: 27413249
- PMCID: PMC4930817
- DOI: 10.1155/2016/7919534
Sleep Deprivation Influences Circadian Gene Expression in the Lateral Habenula
Abstract
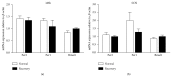
Sleep is governed by homeostasis and the circadian clock. Clock genes play an important role in the generation and maintenance of circadian rhythms but are also involved in regulating sleep homeostasis. The lateral habenular nucleus (LHb) has been implicated in sleep-wake regulation, since LHb gene expression demonstrates circadian oscillation characteristics. This study focuses on the participation of LHb clock genes in regulating sleep homeostasis, as the nature of their involvement is unclear. In this study, we observed changes in sleep pattern following sleep deprivation in LHb-lesioned rats using EEG recording techniques. And then the changes of clock gene expression (Per1, Per2, and Bmal1) in the LHb after 6 hours of sleep deprivation were detected by using real-time quantitative PCR (qPCR). We found that sleep deprivation increased the length of Non-Rapid Eye Movement Sleep (NREMS) and decreased wakefulness. LHb-lesioning decreased the amplitude of reduced wake time and increased NREMS following sleep deprivation in rats. qPCR results demonstrated that Per2 expression was elevated after sleep deprivation, while the other two genes were unaffected. Following sleep recovery, Per2 expression was comparable to the control group. This study provides the basis for further research on the role of LHb Per2 gene in the regulation of sleep homeostasis.
Figures



Similar articles
-
The sleep-wake distribution contributes to the peripheral rhythms in PERIOD-2.Elife. 2021 Dec 13;10:e69773. doi: 10.7554/eLife.69773. Elife. 2021. PMID: 34895464 Free PMC article.
-
Diurnal Expression of the Per2 Gene and Protein in the Lateral Habenular Nucleus.Int J Mol Sci. 2015 Jul 23;16(8):16740-9. doi: 10.3390/ijms160816740. Int J Mol Sci. 2015. PMID: 26213916 Free PMC article.
-
Depression-like behavior is associated with lower Per2 mRNA expression in the lateral habenula of rats.Genes Brain Behav. 2021 Apr;20(4):e12702. doi: 10.1111/gbb.12702. Epub 2020 Oct 12. Genes Brain Behav. 2021. PMID: 32964673
-
Mechanisms of rapid antidepressant effects of sleep deprivation therapy: clock genes and circadian rhythms.Biol Psychiatry. 2013 Jun 15;73(12):1164-71. doi: 10.1016/j.biopsych.2012.07.020. Epub 2012 Aug 18. Biol Psychiatry. 2013. PMID: 22906517 Review.
-
Circadian neurons in the lateral habenula: Clocking motivated behaviors.Pharmacol Biochem Behav. 2017 Nov;162:55-61. doi: 10.1016/j.pbb.2017.06.013. Epub 2017 Jun 28. Pharmacol Biochem Behav. 2017. PMID: 28666896 Review.
Cited by
-
The role of clock genes in sleep, stress and memory.Biochem Pharmacol. 2021 Sep;191:114493. doi: 10.1016/j.bcp.2021.114493. Epub 2021 Feb 27. Biochem Pharmacol. 2021. PMID: 33647263 Free PMC article. Review.
-
A Pattern to Link Adenosine Signaling, Circadian System, and Potential Final Common Pathway in the Pathogenesis of Major Depressive Disorder.Mol Neurobiol. 2022 Nov;59(11):6713-6723. doi: 10.1007/s12035-022-03001-3. Epub 2022 Aug 23. Mol Neurobiol. 2022. PMID: 35999325 Free PMC article. Review.
-
Identification of hub genes correlated with sleep deprivation using co-expression analysis.Sleep Breath. 2021 Dec;25(4):1969-1976. doi: 10.1007/s11325-021-02321-3. Epub 2021 Feb 22. Sleep Breath. 2021. PMID: 33619665
-
Glymphatic Dysfunction: A Bridge Between Sleep Disturbance and Mood Disorders.Front Psychiatry. 2021 May 7;12:658340. doi: 10.3389/fpsyt.2021.658340. eCollection 2021. Front Psychiatry. 2021. PMID: 34025481 Free PMC article.
-
The sleep-wake distribution contributes to the peripheral rhythms in PERIOD-2.Elife. 2021 Dec 13;10:e69773. doi: 10.7554/eLife.69773. Elife. 2021. PMID: 34895464 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

