Densely ionizing radiation affects DNA methylation of selective LINE-1 elements
- PMID: 27419368
- PMCID: PMC5003736
- DOI: 10.1016/j.envres.2016.06.043
Densely ionizing radiation affects DNA methylation of selective LINE-1 elements
Abstract
Long Interspersed Nucleotide Element 1 (LINE-1) retrotransposons are heavily methylated and are the most abundant transposable elements in mammalian genomes. Here, we investigated the differential DNA methylation within the LINE-1 under normal conditions and in response to environmentally relevant doses of sparsely and densely ionizing radiation. We demonstrate that DNA methylation of LINE-1 elements in the lungs of C57BL6 mice is dependent on their evolutionary age, where the elder age of the element is associated with the lower extent of DNA methylation. Exposure to 5-aza-2'-deoxycytidine and methionine-deficient diet affected DNA methylation of selective LINE-1 elements in an age- and promoter type-dependent manner. Exposure to densely IR, but not sparsely IR, resulted in DNA hypermethylation of older LINE-1 elements, while the DNA methylation of evolutionary younger elements remained mostly unchanged. We also demonstrate that exposure to densely IR increased mRNA and protein levels of LINE-1 via the loss of the histone H3K9 dimethylation and an increase in the H3K4 trimethylation at the LINE-1 5'-untranslated region, independently of DNA methylation. Our findings suggest that DNA methylation is important for regulation of LINE-1 expression under normal conditions, but histone modifications may dictate the transcriptional activity of LINE-1 in response to exposure to densely IR.
Keywords: Epigenetics; High-LET radiation; Histone methylation; Ionizing radiation; Transposable elements.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures

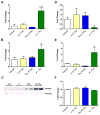
Comment in
-
LINE-1 in response to exposure to ionizing radiation.Mob Genet Elements. 2017 Oct 19;7(6):e1393491. doi: 10.1080/2159256X.2017.1393491. eCollection 2017. Mob Genet Elements. 2017. PMID: 29209557 Free PMC article.
Similar articles
-
Changes in one-carbon metabolism and DNA methylation in the hearts of mice exposed to space environment-relevant doses of oxygen ions (16O).Life Sci Space Res (Amst). 2019 Aug;22:8-15. doi: 10.1016/j.lssr.2019.05.003. Epub 2019 May 31. Life Sci Space Res (Amst). 2019. PMID: 31421852 Free PMC article.
-
Inter-Strain Differences in LINE-1 DNA Methylation in the Mouse Hematopoietic System in Response to Exposure to Ionizing Radiation.Int J Mol Sci. 2017 Jul 4;18(7):1430. doi: 10.3390/ijms18071430. Int J Mol Sci. 2017. PMID: 28677663 Free PMC article.
-
Radiation-induced changes in DNA methylation of repetitive elements in the mouse heart.Mutat Res. 2016 May;787:43-53. doi: 10.1016/j.mrfmmm.2016.02.009. Epub 2016 Mar 2. Mutat Res. 2016. PMID: 26963372 Free PMC article.
-
LINEs in mice: features, families, and potential roles in early development.Chromosoma. 2016 Mar;125(1):29-39. doi: 10.1007/s00412-015-0520-2. Epub 2015 May 16. Chromosoma. 2016. PMID: 25975894 Review.
-
Long Interspersed Element-1 Methylation Level as a Prognostic Biomarker in Gastrointestinal Cancers.Digestion. 2018;97(1):26-30. doi: 10.1159/000484104. Epub 2018 Feb 1. Digestion. 2018. PMID: 29393154 Review.
Cited by
-
Irradiation and Alterations in Hippocampal DNA Methylation.Epigenomes. 2024 Jul 5;8(3):27. doi: 10.3390/epigenomes8030027. Epigenomes. 2024. PMID: 39051185 Free PMC article. Review.
-
Ionizing radiation induces transgenerational effects of DNA methylation in zebrafish.Sci Rep. 2018 Oct 18;8(1):15373. doi: 10.1038/s41598-018-33817-w. Sci Rep. 2018. PMID: 30337673 Free PMC article.
-
Changes in one-carbon metabolism and DNA methylation in the hearts of mice exposed to space environment-relevant doses of oxygen ions (16O).Life Sci Space Res (Amst). 2019 Aug;22:8-15. doi: 10.1016/j.lssr.2019.05.003. Epub 2019 May 31. Life Sci Space Res (Amst). 2019. PMID: 31421852 Free PMC article.
-
Climate Change and New Challenges for Rural Communities: Particulate Matter Matters.Sustainability. 2023 Dec 1;15(23):16192. doi: 10.3390/su152316192. Epub 2023 Nov 22. Sustainability. 2023. PMID: 39119507 Free PMC article.
-
Effects of ionizing radiation on DNA methylation: from experimental biology to clinical applications.Int J Radiat Biol. 2017 May;93(5):457-469. doi: 10.1080/09553002.2017.1287454. Epub 2017 Feb 21. Int J Radiat Biol. 2017. PMID: 28134023 Free PMC article. Review.
References
-
- Brooks A, Bao S, Rithidech K, Couch LA, Braby LA. Relative effectiveness of HZE iron-56 particles for the induction of cytogenetic damage in vivo. Radiat Res. 2001;155:353–359. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

