AcmB Is an S-Layer-Associated β-N-Acetylglucosaminidase and Functional Autolysin in Lactobacillus acidophilus NCFM
- PMID: 27422832
- PMCID: PMC5007774
- DOI: 10.1128/AEM.02025-16
AcmB Is an S-Layer-Associated β-N-Acetylglucosaminidase and Functional Autolysin in Lactobacillus acidophilus NCFM
Abstract
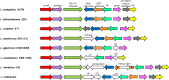
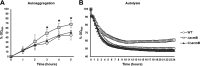
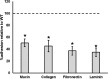
Autolysins, also known as peptidoglycan hydrolases, are enzymes that hydrolyze specific bonds within bacterial cell wall peptidoglycan during cell division and daughter cell separation. Within the genome of Lactobacillus acidophilus NCFM, there are 11 genes encoding proteins with peptidoglycan hydrolase catalytic domains, 9 of which are predicted to be functional. Notably, 5 of the 9 putative autolysins in L. acidophilus NCFM are S-layer-associated proteins (SLAPs) noncovalently colocalized along with the surface (S)-layer at the cell surface. One of these SLAPs, AcmB, a β-N-acetylglucosaminidase encoded by the gene lba0176 (acmB), was selected for functional analysis. In silico analysis revealed that acmB orthologs are found exclusively in S-layer- forming species of Lactobacillus Chromosomal deletion of acmB resulted in aberrant cell division, autolysis, and autoaggregation. Complementation of acmB in the ΔacmB mutant restored the wild-type phenotype, confirming the role of this SLAP in cell division. The absence of AcmB within the exoproteome had a pleiotropic effect on the extracellular proteins covalently and noncovalently bound to the peptidoglycan, which likely led to the observed decrease in the binding capacity of the ΔacmB strain for mucin and extracellular matrices fibronectin, laminin, and collagen in vitro These data suggest a functional association between the S-layer and the multiple autolysins noncovalently colocalized at the cell surface of L. acidophilus NCFM and other S-layer-producing Lactobacillus species.
Importance: Lactobacillus acidophilus is one of the most widely used probiotic microbes incorporated in many dairy foods and dietary supplements. This organism produces a surface (S)-layer, which is a self-assembling crystalline array found as the outermost layer of the cell wall. The S-layer, along with colocalized associated proteins, is an important mediator of probiotic activity through intestinal adhesion and modulation of the mucosal immune system. However, there is still a dearth of information regarding the basic cellular and evolutionary function of S-layers. Here, we demonstrate that multiple autolysins, responsible for breaking down the cell wall during cell division, are associated with the S-layer. Deletion of the gene encoding one of these S-layer-associated autolysins confirmed its autolytic role and resulted in reduced binding capacity to mucin and intestinal extracellular matrices. These data suggest a functional association between the S-layer and autolytic activity through the extracellular presentation of autolysins.
Copyright © 2016 Johnson and Klaenhammer.
Figures

Similar articles
-
Functional Analysis of an S-Layer-Associated Fibronectin-Binding Protein in Lactobacillus acidophilus NCFM.Appl Environ Microbiol. 2016 Apr 18;82(9):2676-2685. doi: 10.1128/AEM.00024-16. Print 2016 May. Appl Environ Microbiol. 2016. PMID: 26921419 Free PMC article.
-
Characterization of AcmB, an N-acetylglucosaminidase autolysin from Lactococcus lactis.Microbiology (Reading). 2003 Mar;149(Pt 3):695-705. doi: 10.1099/mic.0.25875-0. Microbiology (Reading). 2003. PMID: 12634338
-
Identification of key peptidoglycan hydrolases for morphogenesis, autolysis, and peptidoglycan composition of Lactobacillus plantarum WCFS1.Microb Cell Fact. 2012 Oct 15;11:137. doi: 10.1186/1475-2859-11-137. Microb Cell Fact. 2012. PMID: 23066986 Free PMC article.
-
Invited review: the scientific basis of Lactobacillus acidophilus NCFM functionality as a probiotic.J Dairy Sci. 2001 Feb;84(2):319-31. doi: 10.3168/jds.S0022-0302(01)74481-5. J Dairy Sci. 2001. PMID: 11233016 Review.
-
Bacterial peptidoglycan (murein) hydrolases.FEMS Microbiol Rev. 2008 Mar;32(2):259-86. doi: 10.1111/j.1574-6976.2007.00099.x. Epub 2008 Feb 11. FEMS Microbiol Rev. 2008. PMID: 18266855 Review.
Cited by
-
An Acid Up-Regulated Surface Protein of Lactobacillus paracasei Strain GCRL 46 is Phylogenetically Related to the Secreted Glucan- (GpbB) and Immunoglobulin-Binding (SibA) Protein of Pathogenic Streptococci.Int J Mol Sci. 2019 Mar 31;20(7):1610. doi: 10.3390/ijms20071610. Int J Mol Sci. 2019. PMID: 30935131 Free PMC article.
-
S-layer associated proteins contribute to the adhesive and immunomodulatory properties of Lactobacillus acidophilus NCFM.BMC Microbiol. 2020 Aug 12;20(1):248. doi: 10.1186/s12866-020-01908-2. BMC Microbiol. 2020. PMID: 32787778 Free PMC article.
-
The S-layer Associated Serine Protease Homolog PrtX Impacts Cell Surface-Mediated Microbe-Host Interactions of Lactobacillus acidophilus NCFM.Front Microbiol. 2017 Jun 30;8:1185. doi: 10.3389/fmicb.2017.01185. eCollection 2017. Front Microbiol. 2017. PMID: 28713337 Free PMC article.
-
Probiotic Gastrointestinal Transit and Colonization After Oral Administration: A Long Journey.Front Cell Infect Microbiol. 2021 Mar 10;11:609722. doi: 10.3389/fcimb.2021.609722. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 33791234 Free PMC article. Review.
-
Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes.Int J Mol Sci. 2022 Dec 2;23(23):15144. doi: 10.3390/ijms232315144. Int J Mol Sci. 2022. PMID: 36499471 Free PMC article.
References
-
- Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, Morelli L, Canani RB, Flint HJ, Salminen S, Calder PC, Sanders ME. 2014. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11:506–514. doi:10.1038/nrgastro.2014.66. - DOI - PubMed
-
- Altermann E, Russell WM, Azcarate-Peril MA, Barrangou R, Buck BL, McAuliffe O, Souther N, Dobson A, Duong T, Callanan M, Lick S, Hamrick A, Cano R, Klaenhammer TR. 2005. Complete genome sequence of the probiotic lactic acid bacterium Lactobacillus acidophilus NCFM. Proc Natl Acad Sci U S A 102:3906–3912. doi:10.1073/pnas.0409188102. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

