Upregulated LINE-1 Activity in the Fanconi Anemia Cancer Susceptibility Syndrome Leads to Spontaneous Pro-inflammatory Cytokine Production
- PMID: 27428429
- PMCID: PMC4919473
- DOI: 10.1016/j.ebiom.2016.05.005
Upregulated LINE-1 Activity in the Fanconi Anemia Cancer Susceptibility Syndrome Leads to Spontaneous Pro-inflammatory Cytokine Production
Abstract
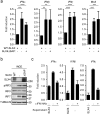
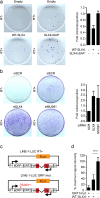
Fanconi Anemia (FA) is a genetic disorder characterized by elevated cancer susceptibility and pro-inflammatory cytokine production. Using SLX4(FANCP) deficiency as a working model, we questioned the trigger for chronic inflammation in FA. We found that absence of SLX4 caused cytoplasmic DNA accumulation, including sequences deriving from active Long INterspersed Element-1 (LINE-1), triggering the cGAS-STING pathway to elicit interferon (IFN) expression. In agreement, absence of SLX4 leads to upregulated LINE-1 retrotransposition. Importantly, similar results were obtained with the FANCD2 upstream activator of SLX4. Furthermore, treatment of FA cells with the Tenofovir reverse transcriptase inhibitor (RTi), that prevents endogenous retrotransposition, decreased both accumulation of cytoplasmic DNA and pro-inflammatory signaling. Collectively, our data suggest a contribution of endogenous RT activities to the generation of immunogenic cytoplasmic nucleic acids responsible for inflammation in FA. The additional observation that RTi decreased pro-inflammatory cytokine production induced by DNA replication stress-inducing drugs further demonstrates the contribution of endogenous RTs to sustaining chronic inflammation. Altogether, our data open perspectives in the prevention of adverse effects of chronic inflammation in tumorigenesis.
Keywords: Cytoplasmic DNA; DNA damage; Fanconi Anemia; Inflammation; Innate immune sensing; Interferon; SLX4 complex; cGAS-STING; endogenous retroelement.
Copyright © 2016 The Authors. Published by Elsevier B.V. All rights reserved.
Figures





Comment in
-
Multifunctional Fanconi proteins, inflammation and the Fanconi phenotype.EBioMedicine. 2016 Jun;8:10-11. doi: 10.1016/j.ebiom.2016.06.005. Epub 2016 Jun 3. EBioMedicine. 2016. PMID: 27428405 Free PMC article. No abstract available.
References
-
- Babushok D.V., Kazazian H.H., Jr. Progress in understanding the biology of the human mutagen LINE-1. Hum. Mutat. 2007;28:527–539. - PubMed
-
- Bald T., Quast T., Landsberg J., Rogava M., Glodde N., Lopez-Ramos D., Kohlmeyer J., Riesenberg S., van den Boorn-Konijnenberg D., Homig-Holzel C. Ultraviolet-radiation-induced inflammation promotes angiotropism and metastasis in melanoma. Nature. 2014;507:109–113. - PubMed
-
- Bogliolo M., Surralles J. Fanconi anemia: a model disease for studies on human genetics and advanced therapeutics. Curr. Opin. Genet. Dev. 2015;33:32–40. - PubMed
-
- Boissinot S., Chevret P., Furano A.V. L1 (LINE-1) retrotransposon evolution and amplification in recent human history. Mol. Biol. Evol. 2000;17:915–928. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

