Bile signalling promotes chronic respiratory infections and antibiotic tolerance
- PMID: 27432520
- PMCID: PMC4949476
- DOI: 10.1038/srep29768
Bile signalling promotes chronic respiratory infections and antibiotic tolerance
Abstract
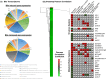
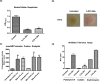
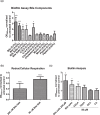
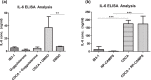
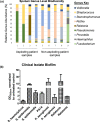
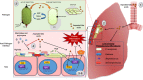
Despite aggressive antimicrobial therapy, many respiratory pathogens persist in the lung, underpinning the chronic inflammation and eventual lung decline that are characteristic of respiratory disease. Recently, bile acid aspiration has emerged as a major comorbidity associated with a range of lung diseases, shaping the lung microbiome and promoting colonisation by Pseudomonas aeruginosa in Cystic Fibrosis (CF) patients. In order to uncover the molecular mechanism through which bile modulates the respiratory microbiome, a combination of global transcriptomic and phenotypic analyses of the P. aeruginosa response to bile was undertaken. Bile responsive pathways responsible for virulence, adaptive metabolism, and redox control were identified, with macrolide and polymyxin antibiotic tolerance increased significantly in the presence of bile. Bile acids, and chenodeoxycholic acid (CDCA) in particular, elicited chronic biofilm behaviour in P. aeruginosa, while induction of the pro-inflammatory cytokine Interleukin-6 (IL-6) in lung epithelial cells by CDCA was Farnesoid X Receptor (FXR) dependent. Microbiome analysis of paediatric CF sputum samples demonstrated increased colonisation by P. aeruginosa and other Proteobacterial pathogens in bile aspirating compared to non-aspirating patients. Together, these data suggest that bile acid signalling is a leading trigger for the development of chronic phenotypes underlying the pathophysiology of chronic respiratory disease.
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

