Silencing of natural transformation by an RNA chaperone and a multitarget small RNA
- PMID: 27432973
- PMCID: PMC4978251
- DOI: 10.1073/pnas.1601626113
Silencing of natural transformation by an RNA chaperone and a multitarget small RNA
Abstract
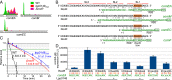
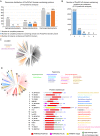
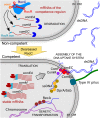
A highly conserved DNA uptake system allows many bacteria to actively import and integrate exogenous DNA. This process, called natural transformation, represents a major mechanism of horizontal gene transfer (HGT) involved in the acquisition of virulence and antibiotic resistance determinants. Despite evidence of HGT and the high level of conservation of the genes coding the DNA uptake system, most bacterial species appear non-transformable under laboratory conditions. In naturally transformable species, the DNA uptake system is only expressed when bacteria enter a physiological state called competence, which develops under specific conditions. Here, we investigated the mechanism that controls expression of the DNA uptake system in the human pathogen Legionella pneumophila We found that a repressor of this system displays a conserved ProQ/FinO domain and interacts with a newly characterized trans-acting sRNA, RocR. Together, they target mRNAs of the genes coding the DNA uptake system to control natural transformation. This RNA-based silencing represents a previously unknown regulatory means to control this major mechanism of HGT. Importantly, these findings also show that chromosome-encoded ProQ/FinO domain-containing proteins can assist trans-acting sRNAs and that this class of RNA chaperones could play key roles in post-transcriptional gene regulation throughout bacterial species.
Keywords: Legionella pneumophila; ProQ/FinO; RNA chaperone; natural transformation; non-coding RNA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Johnston C, Martin B, Fichant G, Polard P, Claverys J-P. Bacterial transformation: Distribution, shared mechanisms and divergent control. Nat Rev Microbiol. 2014;12(3):181–196. - PubMed
-
- Smith JM, Dowson CG, Spratt BG. Localized sex in bacteria. Nature. 1991;349(6304):29–31. - PubMed
-
- Seitz P, Blokesch M. Cues and regulatory pathways involved in natural competence and transformation in pathogenic and environmental Gram-negative bacteria. FEMS Microbiol Rev. 2013;37(3):336–363. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

