Biochemical Characterization of a Family 15 Carbohydrate Esterase from a Bacterial Marine Arctic Metagenome
- PMID: 27433797
- PMCID: PMC4951047
- DOI: 10.1371/journal.pone.0159345
Biochemical Characterization of a Family 15 Carbohydrate Esterase from a Bacterial Marine Arctic Metagenome
Abstract
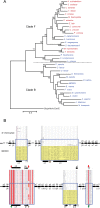
Background: The glucuronoyl esterase enzymes of wood-degrading fungi (Carbohydrate Esterase family 15; CE15) form part of the hemicellulolytic and cellulolytic enzyme systems that break down plant biomass, and have possible applications in biotechnology. Homologous enzymes are predicted in the genomes of several bacteria, however these have been much less studied than their fungal counterparts. Here we describe the recombinant production and biochemical characterization of a bacterial CE15 enzyme denoted MZ0003, which was identified by in silico screening of a prokaryotic metagenome library derived from marine Arctic sediment. MZ0003 has high similarity to several uncharacterized gene products of polysaccharide-degrading bacterial species, and phylogenetic analysis indicates a deep evolutionary split between these CE15s and fungal homologs.
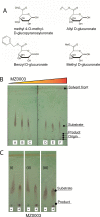
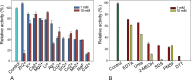
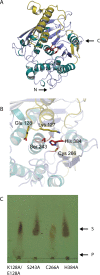
Results: MZ0003 appears to differ from previously-studied CE15s in some aspects. Some glucuronoyl esterase activity could be measured by qualitative thin-layer chromatography which confirms its assignment as a CE15, however MZ0003 can also hydrolyze a range of other esters, including p-nitrophenyl acetate, which is not acted upon by some fungal homologs. The structure of MZ0003 also appears to differ as it is predicted to have several large loop regions that are absent in previously studied CE15s, and a combination of homology-based modelling and site-directed mutagenesis indicate its catalytic residues deviate from the conserved Ser-His-Glu triad of many fungal CE15s. Taken together, these results indicate that potentially unexplored diversity exists among bacterial CE15s, and this may be accessed by investigation of the microbial metagenome. The combination of low activity on typical glucuronoyl esterase substrates, and the lack of glucuronic acid esters in the marine environment suggest that the physiological substrate of MZ0003 and its homologs is likely to be different from that of related fungal enzymes.
Conflict of interest statement
Figures



Similar articles
-
Structural insight into a CE15 esterase from the marine bacterial metagenome.Sci Rep. 2017 Dec 8;7(1):17278. doi: 10.1038/s41598-017-17677-4. Sci Rep. 2017. PMID: 29222424 Free PMC article.
-
Microbial Glucuronoyl Esterases: 10 Years after Discovery.Appl Environ Microbiol. 2016 Nov 21;82(24):7014-7018. doi: 10.1128/AEM.02396-16. Print 2016 Dec 15. Appl Environ Microbiol. 2016. PMID: 27694239 Free PMC article. Review.
-
Structural and biochemical studies of the glucuronoyl esterase OtCE15A illuminate its interaction with lignocellulosic components.J Biol Chem. 2019 Dec 27;294(52):19978-19987. doi: 10.1074/jbc.RA119.011435. Epub 2019 Nov 18. J Biol Chem. 2019. PMID: 31740581 Free PMC article.
-
Identification of genes encoding microbial glucuronoyl esterases.FEBS Lett. 2007 Aug 21;581(21):4029-35. doi: 10.1016/j.febslet.2007.07.041. Epub 2007 Jul 26. FEBS Lett. 2007. PMID: 17678650
-
Glucuronoyl esterases: diversity, properties and biotechnological potential. A review.Crit Rev Biotechnol. 2018 Nov;38(7):1121-1136. doi: 10.1080/07388551.2018.1468316. Epub 2018 May 8. Crit Rev Biotechnol. 2018. PMID: 29739247 Review.
Cited by
-
Structure-function analyses reveal that a glucuronoyl esterase from Teredinibacter turnerae interacts with carbohydrates and aromatic compounds.J Biol Chem. 2019 Apr 19;294(16):6635-6644. doi: 10.1074/jbc.RA119.007831. Epub 2019 Feb 27. J Biol Chem. 2019. PMID: 30814248 Free PMC article.
-
Biochemical and structural features of diverse bacterial glucuronoyl esterases facilitating recalcitrant biomass conversion.Biotechnol Biofuels. 2018 Aug 1;11:213. doi: 10.1186/s13068-018-1213-x. eCollection 2018. Biotechnol Biofuels. 2018. PMID: 30083226 Free PMC article.
-
Structural insight into a CE15 esterase from the marine bacterial metagenome.Sci Rep. 2017 Dec 8;7(1):17278. doi: 10.1038/s41598-017-17677-4. Sci Rep. 2017. PMID: 29222424 Free PMC article.
-
New insights to diversity and enzyme-substrate interactions of fungal glucuronoyl esterases.Appl Microbiol Biotechnol. 2023 Jul;107(14):4447-4457. doi: 10.1007/s00253-023-12575-4. Epub 2023 May 31. Appl Microbiol Biotechnol. 2023. PMID: 37256329 Free PMC article. Review.
-
Mechanism and biomass association of glucuronoyl esterase: an α/β hydrolase with potential in biomass conversion.Nat Commun. 2022 Mar 18;13(1):1449. doi: 10.1038/s41467-022-28938-w. Nat Commun. 2022. PMID: 35304453 Free PMC article.
References
-
- Spanikova S, Biely P. Glucuronoyl esterase-novel carbohydrate esterase produced by Schizophyllum commune. Febs Lett. 2006;580(19):4597–601. - PubMed
-
- Spanikova S, Polakova M, Joniak D, Hirsch J, Biely P. Synthetic esters recognized by glucuronoyl esterase from Schizophyllum commune. Arch Microbiol. 2007;188(2):185–9. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

