Outer membrane protein functions as integrator of protein import and DNA inheritance in mitochondria
- PMID: 27436903
- PMCID: PMC4978248
- DOI: 10.1073/pnas.1605497113
Outer membrane protein functions as integrator of protein import and DNA inheritance in mitochondria
Abstract
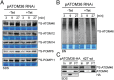
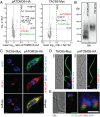
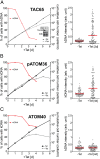
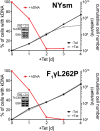
Trypanosomatids are one of the earliest diverging eukaryotes that have fully functional mitochondria. pATOM36 is a trypanosomatid-specific essential mitochondrial outer membrane protein that has been implicated in protein import. Changes in the mitochondrial proteome induced by ablation of pATOM36 and in vitro assays show that pATOM36 is required for the assembly of the archaic translocase of the outer membrane (ATOM), the functional analog of the TOM complex in other organisms. Reciprocal pull-down experiments and immunofluorescence analyses demonstrate that a fraction of pATOM36 interacts and colocalizes with TAC65, a previously uncharacterized essential component of the tripartite attachment complex (TAC). The TAC links the single-unit mitochondrial genome to the basal body of the flagellum and mediates the segregation of the replicated mitochondrial genomes. RNAi experiments show that pATOM36, in line with its dual localization, is not only essential for ATOM complex assembly but also for segregation of the replicated mitochondrial genomes. However, the two functions are distinct, as a truncated version of pATOM36 lacking the 75 C-terminal amino acids can rescue kinetoplast DNA missegregation but not the lack of ATOM complex assembly. Thus, pATOM36 has a dual function and integrates mitochondrial protein import with mitochondrial DNA inheritance.
Keywords: evolution; mitochondria; mitochondrial genome; protein import; trypanosomes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






Similar articles
-
Independent evolution of functionally exchangeable mitochondrial outer membrane import complexes.Elife. 2018 Jun 20;7:e34488. doi: 10.7554/eLife.34488. Elife. 2018. PMID: 29923829 Free PMC article.
-
An essential novel component of the noncanonical mitochondrial outer membrane protein import system of trypanosomatids.Mol Biol Cell. 2012 Sep;23(17):3420-8. doi: 10.1091/mbc.E12-02-0107. Epub 2012 Jul 11. Mol Biol Cell. 2012. PMID: 22787278 Free PMC article.
-
TbLOK1/ATOM19 is a novel subunit of the noncanonical mitochondrial outer membrane protein translocase of Trypanosoma brucei.Mol Microbiol. 2016 Nov;102(3):520-529. doi: 10.1111/mmi.13476. Epub 2016 Aug 18. Mol Microbiol. 2016. PMID: 27501349
-
Mitochondrial protein import in trypanosomes: Expect the unexpected.Traffic. 2017 Feb;18(2):96-109. doi: 10.1111/tra.12463. Epub 2017 Jan 17. Traffic. 2017. PMID: 27976830 Review.
-
Failure is not an option - mitochondrial genome segregation in trypanosomes.J Cell Sci. 2018 Sep 17;131(18):jcs221820. doi: 10.1242/jcs.221820. J Cell Sci. 2018. PMID: 30224426 Review.
Cited by
-
A comparison of three approaches for the discovery of novel tripartite attachment complex proteins in Trypanosoma brucei.PLoS Negl Trop Dis. 2020 Sep 16;14(9):e0008568. doi: 10.1371/journal.pntd.0008568. eCollection 2020 Sep. PLoS Negl Trop Dis. 2020. PMID: 32936798 Free PMC article.
-
Independent evolution of functionally exchangeable mitochondrial outer membrane import complexes.Elife. 2018 Jun 20;7:e34488. doi: 10.7554/eLife.34488. Elife. 2018. PMID: 29923829 Free PMC article.
-
Down the membrane hole: Ion channels in protozoan parasites.PLoS Pathog. 2022 Dec 29;18(12):e1011004. doi: 10.1371/journal.ppat.1011004. eCollection 2022 Dec. PLoS Pathog. 2022. PMID: 36580479 Free PMC article. Review.
-
Mitochondrial protein import in trypanosomatids: Variations on a theme or fundamentally different?PLoS Pathog. 2018 Nov 29;14(11):e1007351. doi: 10.1371/journal.ppat.1007351. eCollection 2018 Nov. PLoS Pathog. 2018. PMID: 30496284 Free PMC article. Review. No abstract available.
-
p166 links membrane and intramitochondrial modules of the trypanosomal tripartite attachment complex.PLoS Pathog. 2022 Jun 16;18(6):e1010207. doi: 10.1371/journal.ppat.1010207. eCollection 2022 Jun. PLoS Pathog. 2022. PMID: 35709300 Free PMC article.
References
-
- Westermann B. Mitochondrial inheritance in yeast. Biochim Biophys Acta. 2014;1837(7):1039–1046. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

