A network of genes associated with poplar root development in response to low nitrogen
- PMID: 27449227
- PMCID: PMC5022414
- DOI: 10.1080/15592324.2016.1214792
A network of genes associated with poplar root development in response to low nitrogen
Abstract
Deployment of the root system is highly sensitive to the levels and spatial distribution of nutrients like nitrogen. However, the genetic determinants of these sensing and deployment mechanisms are still poorly understood. Previously, using system approaches based on temporal changes in root transcriptome in relation to low nitrogen (LN), we have been able to identify a module that activates root production in poplar in response to LN conditions. Here, using comparative, gene ontology and expression analyses, we provide further evidence that the genes in this module are indeed involved in regulation of root development under LN. Better understanding of these modules will enable approaches for breeding for better nitrogen use efficiency through development of a more sensitive and plastic root system.
Keywords: Gene network; Populus tremula x P. alba; hubs; lateral root growth; nitrogen stress.
Figures

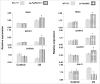
Similar articles
-
A systems biology approach identifies new regulators of poplar root development under low nitrogen.Plant J. 2015 Oct;84(2):335-46. doi: 10.1111/tpj.13002. Plant J. 2015. PMID: 26315649
-
Nitrogen deprivation promotes Populus root growth through global transcriptome reprogramming and activation of hierarchical genetic networks.New Phytol. 2013 Oct;200(2):483-497. doi: 10.1111/nph.12375. Epub 2013 Jun 25. New Phytol. 2013. PMID: 23795675
-
Interaction of nitrogen nutrition and salinity in Grey poplar (Populus tremula x alba).Plant Cell Environ. 2007 Jul;30(7):796-811. doi: 10.1111/j.1365-3040.2007.01668.x. Plant Cell Environ. 2007. PMID: 17547652
-
Root uptake regulation: a central process for NPS homeostasis in plants.Curr Opin Plant Biol. 2009 Jun;12(3):328-38. doi: 10.1016/j.pbi.2009.04.015. Epub 2009 Jun 6. Curr Opin Plant Biol. 2009. PMID: 19501015 Review.
-
Demystifying roots: A need for clarification and extended concepts in root phenotyping.Plant Sci. 2019 May;282:11-13. doi: 10.1016/j.plantsci.2018.09.015. Epub 2018 Sep 25. Plant Sci. 2019. PMID: 31003606 Review.
Cited by
-
Influence of Nitrogen Availability on Growth of Two Transgenic Birch Species Carrying the Pine GS1a Gene.Plants (Basel). 2017 Jan 6;6(1):4. doi: 10.3390/plants6010004. Plants (Basel). 2017. PMID: 28067821 Free PMC article.
-
Genome-Wide Identification of WRKY Family Genes and the Expression Profiles in Response to Nitrogen Deficiency in Poplar.Genes (Basel). 2022 Dec 10;13(12):2324. doi: 10.3390/genes13122324. Genes (Basel). 2022. PMID: 36553591 Free PMC article.
-
Combining GS-assisted GWAS and transcriptome analysis to mine candidate genes for nitrogen utilization efficiency in Populus cathayana.BMC Plant Biol. 2023 Apr 5;23(1):182. doi: 10.1186/s12870-023-04202-1. BMC Plant Biol. 2023. PMID: 37020197 Free PMC article.
-
Morphological, physiological, and transcriptional responses to low nitrogen stress in Populus deltoides Marsh. clones with contrasting nitrogen use efficiency.BMC Genomics. 2021 Sep 27;22(1):697. doi: 10.1186/s12864-021-07991-7. BMC Genomics. 2021. PMID: 34579659 Free PMC article.
-
Identification and Functional Prediction of Poplar Root circRNAs Involved in Treatment With Different Forms of Nitrogen.Front Plant Sci. 2022 Jul 8;13:941380. doi: 10.3389/fpls.2022.941380. eCollection 2022. Front Plant Sci. 2022. PMID: 35874008 Free PMC article.
References
-
- Dash M, Yordanov YS, Georgieva T, Kumari S, Wei H, Busov V. A systems biology approach identifies new regulators of poplar root development under low nitrogen. Plant J 2015; 84:335-46; PMID:26315649; http://dx.doi.org/10.1111/tpj.13002 - DOI - PubMed
-
- Wei HR, Yordanov YS, Georgieva T, Li X, Busov V. Nitrogen deprivation promotes Populus root growth through global transcriptome reprogramming and activation of hierarchical genetic networks. New Phytol 2013; 200:483-97; PMID:23795675; http://dx.doi.org/10.1111/nph.12375 - DOI - PubMed
-
- Wei H, Yordanov YS, Kumari S, Georgieva T, Busov V. Genetic networks involved in poplar root response to low nitrogen. Plant Signal Behav 2013; 8:483-97; PMID:24300216; http://dx.doi.org/10.4161/psb.27211 - DOI - PMC - PubMed
-
- Lee HW, Kim J. EXPANSINA17 up-regulated by LBD18/ASL20 promotes lateral root formation during the auxin response. Plant Cell Physiol 2013:pct105; PMID:23872272; http://dx.doi.org/1590860010.1093/pcp/pct105 - DOI - PubMed
-
- Yuen CY, Sedbrook JC, Perrin RM, Carroll KL, Masson PH. Loss-of-function mutations of ROOT HAIR DEFECTIVE3 suppress root waving, skewing, and epidermal cell file rotation in Arabidopsis. Plant Physiol 2005; 138:701-14; PMID:15908600; http://dx.doi.org/10.1104/pp.105.059774 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
