Altered Co-Translational Processing Plays a Role in Huntington's Pathogenesis-A Hypothesis
- PMID: 27458341
- PMCID: PMC4933702
- DOI: 10.3389/fnmol.2016.00054
Altered Co-Translational Processing Plays a Role in Huntington's Pathogenesis-A Hypothesis
Abstract
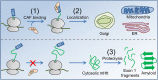
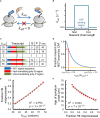
Huntington's disease (HD) is an autosomal dominant neurodegenerative disorder caused by the expansion of a CAG codon repeat region in the HTT gene's first exon that results in huntingtin protein aggregation and neuronal cell death. The development of therapeutic treatments for HD is hindered by the fact that while the etiology and symptoms of HD are understood, the molecular processes connecting this genotype to its phenotype remain unclear. Here, we propose the novel hypothesis that the perturbation of a co-translational process affects mutant huntingtin due to altered translation-elongation kinetics. These altered kinetics arise from the shift of a proline-induced translational pause site away from Htt's localization sequence due to the expansion of the CAG-repeat segment between the poly-proline and localization sequences. Motivation for this hypothesis comes from recent experiments in the field of protein biogenesis that illustrate the critical role that temporal coordination of co-translational processes plays in determining the function, localization, and fate of proteins in cells. We show that our hypothesis is consistent with various experimental observations concerning HD pathology, including the dependence of the age of symptom onset on CAG repeat number. Finally, we suggest three experiments to test our hypothesis.
Keywords: Huntington's disease; biophysics; kinetics; neurodegenerative disease; protein aggregation; protein biogenesis; translation; translation regulation.
Figures
Similar articles
-
Huntington's Disease: Relationship Between Phenotype and Genotype.Mol Neurobiol. 2017 Jan;54(1):342-348. doi: 10.1007/s12035-015-9662-8. Epub 2016 Jan 7. Mol Neurobiol. 2017. PMID: 26742514 Review.
-
Pathogenesis of Huntington's Disease: An Emphasis on Molecular Pathways and Prevention by Natural Remedies.Brain Sci. 2022 Oct 14;12(10):1389. doi: 10.3390/brainsci12101389. Brain Sci. 2022. PMID: 36291322 Free PMC article. Review.
-
Increased Steady-State Mutant Huntingtin mRNA in Huntington's Disease Brain.J Huntingtons Dis. 2013;2(4):491-500. doi: 10.3233/JHD-130079. J Huntingtons Dis. 2013. PMID: 25062733
-
Sirtuins as Modifiers of Huntington's Disease (HD) Pathology.Prog Mol Biol Transl Sci. 2018;154:105-145. doi: 10.1016/bs.pmbts.2017.11.013. Epub 2017 Dec 27. Prog Mol Biol Transl Sci. 2018. PMID: 29413175 Review.
-
SUMO-modifying Huntington's disease.IBRO Neurosci Rep. 2022 Mar 9;12:203-209. doi: 10.1016/j.ibneur.2022.03.002. eCollection 2022 Jun. IBRO Neurosci Rep. 2022. PMID: 35746980 Free PMC article. Review.
Cited by
-
Identification of contributing genes of Huntington's disease by machine learning.BMC Med Genomics. 2020 Nov 23;13(1):176. doi: 10.1186/s12920-020-00822-w. BMC Med Genomics. 2020. PMID: 33228685 Free PMC article.
-
Disome-seq reveals widespread ribosome collisions that promote cotranslational protein folding.Genome Biol. 2021 Jan 5;22(1):16. doi: 10.1186/s13059-020-02256-0. Genome Biol. 2021. PMID: 33402206 Free PMC article.
References
-
- Bartoszewski R. A., Jablonsky M., Bartoszewska S., Stevenson L., Dai Q., Kappes J., et al. . (2010). A synonymous single nucleotide polymorphism in DeltaF508 CFTR alters the secondary structure of the mRNA and the expression of the mutant protein. J. Biol. Chem. 285, 28741–28748. 10.1074/jbc.M110.154575 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

