eIF2 interactions with initiator tRNA and eIF2B are regulated by post-translational modifications and conformational dynamics
- PMID: 27462419
- PMCID: PMC4860841
- DOI: 10.1038/celldisc.2015.20
eIF2 interactions with initiator tRNA and eIF2B are regulated by post-translational modifications and conformational dynamics
Abstract
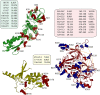
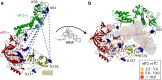
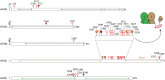
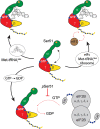
Translation of messenger RNA (mRNA) into proteins is key to eukaryotic gene expression and begins when initiation factor-2 (eIF2) delivers methionyl initiator tRNA (Met-tRNAi (Met)) to ribosomes. This first step is controlled by eIF2B mediating guanine nucleotide exchange on eIF2. We isolated eIF2 from yeast and used mass spectrometry to study the intact complex, and found that eIF2β is the most labile of the three subunits (eIF2α/β/γ). We then compared conformational dynamics of the ternary complex eIF2:GTP:Met-tRNAi (Met) with apo eIF2 using comparative chemical cross-linking. Results revealed high conformational dynamics for eIF2α in apo eIF2 while in the ternary complex all three subunits are constrained. Novel post-translational modifications identified here in both eIF2 and eIF2B were combined with established sites, and located within protein sequences and homology models. We found clustering at subunit interfaces and highly phosphorylated unstructured regions, at the N-terminus of eIF2β, and also between the eIF2Bε core and catalytic domains. We propose that modifications of these unstructured regions have a key role in regulating interactions between eIF2 and eIF2B, as well as other eIFs.
Keywords: acetylation; eukaryotic translation initiation; mass spectrometry; phosphorylation.
Figures


Similar articles
-
Tight binding of the phosphorylated alpha subunit of initiation factor 2 (eIF2alpha) to the regulatory subunits of guanine nucleotide exchange factor eIF2B is required for inhibition of translation initiation.Mol Cell Biol. 2001 Aug;21(15):5018-30. doi: 10.1128/MCB.21.15.5018-5030.2001. Mol Cell Biol. 2001. PMID: 11438658 Free PMC article.
-
Identification of domains and residues within the epsilon subunit of eukaryotic translation initiation factor 2B (eIF2Bepsilon) required for guanine nucleotide exchange reveals a novel activation function promoted by eIF2B complex formation.Mol Cell Biol. 2000 Jun;20(11):3965-76. doi: 10.1128/MCB.20.11.3965-3976.2000. Mol Cell Biol. 2000. PMID: 10805739 Free PMC article.
-
Crystal structure of eIF2B and insights into eIF2-eIF2B interactions.FEBS J. 2017 Mar;284(6):868-874. doi: 10.1111/febs.13896. Epub 2016 Sep 29. FEBS J. 2017. PMID: 27627185 Review.
-
Biochemical analysis of the eIF2beta gamma complex reveals a structural function for eIF2alpha in catalyzed nucleotide exchange.J Biol Chem. 2001 Jan 12;276(2):1051-6. doi: 10.1074/jbc.M007398200. J Biol Chem. 2001. PMID: 11042214
-
Clues to the mechanism of action of eIF2B, the guanine-nucleotide-exchange factor for translation initiation.Biochem Soc Trans. 2008 Aug;36(Pt 4):658-64. doi: 10.1042/BST0360658. Biochem Soc Trans. 2008. PMID: 18631136 Review.
Cited by
-
Regulation and function of elF2B in neurological and metabolic disorders.Biosci Rep. 2022 Jun 30;42(6):BSR20211699. doi: 10.1042/BSR20211699. Biosci Rep. 2022. PMID: 35579296 Free PMC article. Review.
-
Recruitment of trimeric eIF2 by phosphatase non-catalytic subunit PPP1R15B.Mol Cell. 2024 Feb 1;84(3):506-521.e11. doi: 10.1016/j.molcel.2023.12.011. Epub 2023 Dec 29. Mol Cell. 2024. PMID: 38159565 Free PMC article.
-
In-silico characterization and expression study of eIF genes associated with abiotic stresses in potato (Solanum tuberosum L.).Sci Rep. 2025 Jul 5;15(1):24082. doi: 10.1038/s41598-025-09429-6. Sci Rep. 2025. PMID: 40617866 Free PMC article.
-
Crosslinking mass spectrometry: A link between structural biology and systems biology.Protein Sci. 2021 Apr;30(4):773-784. doi: 10.1002/pro.4045. Epub 2021 Mar 6. Protein Sci. 2021. PMID: 33594738 Free PMC article. Review.
-
Acetylation and phosphorylation control both local and global stability of the chloroplast F1 ATP synthase.Sci Rep. 2017 Mar 9;7:44068. doi: 10.1038/srep44068. Sci Rep. 2017. PMID: 28276484 Free PMC article.
References
-
- Aitken CE , Lorsch JR . A mechanistic overview of translation initiation in eukaryotes. Nat Struct Mol Biol 2012; 19: 568–576. - PubMed
-
- Maag D , Fekete CA , Gryczynski Z , Lorsch JR . A conformational change in the eukaryotic translation preinitiation complex and release of eIF1 signal recognition of the start codon. Mol Cell 2005; 17: 265–275. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

