Temporal activation of XRCC1-mediated DNA repair is essential for muscle differentiation
- PMID: 27462438
- PMCID: PMC4860966
- DOI: 10.1038/celldisc.2015.41
Temporal activation of XRCC1-mediated DNA repair is essential for muscle differentiation
Abstract
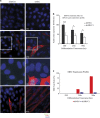
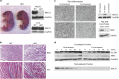
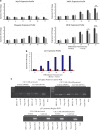
Transient DNA strand break formation has been identified as an effective means to enhance gene expression in living cells. In the muscle lineage, cell differentiation is contingent upon the induction of caspase-mediated DNA strand breaks, which act to establish the terminal gene expression program. This coordinated DNA nicking is rapidly resolved, suggesting that myoblasts may deploy DNA repair machinery to stabilize the genome and entrench the differentiated phenotype. Here, we identify the base excision repair pathway component XRCC1 as an indispensable mediator of muscle differentiation. Caspase-triggered XRCC1 repair foci form rapidly within differentiating myonuclei, and then dissipate as the maturation program proceeds. Skeletal myoblast deletion of Xrcc1 does not have an impact on cell growth, yet leads to perinatal lethality, with sustained DNA damage and impaired myofiber development. Together, these results demonstrate that XRCC1 manages a temporally responsive DNA repair process to advance the muscle differentiation program.
Keywords: DNA strand breaks; XRCC1; base excision repair; muscle differentiation.
Figures


References
-
- Insinga A , Cicalese A , Pelicci PG . DNA damage response in adult stem cells. Blood Cells Mol Dis 2014; 52: 147–151. - PubMed
-
- O’Driscoll M , Jeggo PA . The role of double-strand break repair- insights from human genetics. Nat Rev 2006; 7: 45–54. - PubMed
-
- Roos WP , Kaina B . DNA damage induced cell death: from specific DNA lesions to the DNA damage response and apoptosis. Cancer Lett 2013; 332: 237–248. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

