Tumorigenic potential is restored during differentiation in fusion-reprogrammed cancer cells
- PMID: 27468690
- PMCID: PMC4973342
- DOI: 10.1038/cddis.2016.189
Tumorigenic potential is restored during differentiation in fusion-reprogrammed cancer cells
Abstract
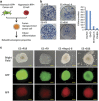
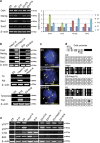
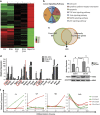
Detailed understanding of the mechanistic steps underlying tumor initiation and malignant progression is critical for insights of potentially novel therapeutic modalities. Cellular reprogramming is an approach of particular interest because it can provide a means to reset the differentiation state of the cancer cells and to revert these cells to a state of non-malignancy. Here, we investigated the relationship between cellular differentiation and malignant progression by the fusion of four independent mouse cancer cell lines from different tissues, each with differing developmental potentials, to pluripotent mouse embryonic stem (ES) cells. Fusion was accompanied by loss of differentiated properties of the four parental cancer cell lines and concomitant emergence of pluripotency, demonstrating the feasibility to reprogram the malignant and differentiative properties of cancer cells. However, the original malignant and differentiative phenotypes re-emerge upon withdrawal of the fused cells from the embryonic environment in which they were maintained. cDNA array analysis of the malignant hepatoma progression implicated a role for Foxa1, and silencing Foxa1 prevented the re-emergence of malignant and differentiation-associated gene expression. Our findings support the hypothesis that tumor progression results from deregulation of stem cells, and our approach provides a strategy to analyze possible mechanisms in the cancer initiation.
Figures



Similar articles
-
Reprogramming of somatic cells after fusion with induced pluripotent stem cells and nuclear transfer embryonic stem cells.Stem Cells Dev. 2010 Feb;19(2):239-46. doi: 10.1089/scd.2009.0142. Stem Cells Dev. 2010. PMID: 19637940
-
Comparison of American mink embryonic stem and induced pluripotent stem cell transcriptomes.BMC Genomics. 2015;16 Suppl 13(Suppl 13):S6. doi: 10.1186/1471-2164-16-S13-S6. Epub 2015 Dec 16. BMC Genomics. 2015. PMID: 26694224 Free PMC article.
-
Generation of induced pluripotent stem cells by efficient reprogramming of adult bone marrow cells.Stem Cells Dev. 2010 Feb;19(2):229-38. doi: 10.1089/scd.2009.0149. Stem Cells Dev. 2010. PMID: 19558219
-
Pluripotent states of human embryonic stem cells.Cell Reprogram. 2015 Feb;17(1):1-6. doi: 10.1089/cell.2014.0061. Epub 2014 Nov 13. Cell Reprogram. 2015. PMID: 25393391 Free PMC article. Review.
-
Teratoma: from spontaneous tumors to the pluripotency/malignancy assay.Wiley Interdiscip Rev Dev Biol. 2016 Mar-Apr;5(2):186-209. doi: 10.1002/wdev.219. Epub 2015 Dec 23. Wiley Interdiscip Rev Dev Biol. 2016. PMID: 26698368 Review.
Cited by
-
Generation of Cancer Stem/Initiating Cells by Cell-Cell Fusion.Int J Mol Sci. 2022 Apr 19;23(9):4514. doi: 10.3390/ijms23094514. Int J Mol Sci. 2022. PMID: 35562905 Free PMC article. Review.
-
Hybrid Formation and Fusion of Cancer Cells In Vitro and In Vivo.Cancers (Basel). 2021 Sep 6;13(17):4496. doi: 10.3390/cancers13174496. Cancers (Basel). 2021. PMID: 34503305 Free PMC article. Review.
-
Detecting the long non‑coding RNA signature related to spinal cord ependymal tumor subtype using a genome‑wide methylome analysis approach.Mol Med Rep. 2019 Aug;20(2):1531-1540. doi: 10.3892/mmr.2019.10388. Epub 2019 Jun 18. Mol Med Rep. 2019. PMID: 31257484 Free PMC article.
-
Deciphering two decades of cellular reprogramming in cancer: A bibliometric analysis of evolving trends and research frontiers.Heliyon. 2024 May 16;10(11):e31400. doi: 10.1016/j.heliyon.2024.e31400. eCollection 2024 Jun 15. Heliyon. 2024. PMID: 38832277 Free PMC article.
-
Nuclear transplantation between allogeneic cells through topological reconnection of plasma membrane in a microfluidic system.Biomicrofluidics. 2019 Jun 10;13(3):034115. doi: 10.1063/1.5098829. eCollection 2019 May. Biomicrofluidics. 2019. PMID: 31312284 Free PMC article.
References
-
- Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature 2004; 429: 457–463. - PubMed
-
- Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev 2002; 3: 415–428. - PubMed
-
- Hahn WC, Weinberg RA. Rules for making human tumor cells. N Engl J Med 2002; 347: 1593–1603. - PubMed
-
- Dougherty GJ, Dougherty ST. Exploiting the tumor microenvironment in the development of targeted cancer gene therapy. Cancer Gene Ther 2009; 16: 279–290. - PubMed
-
- Topczewska JM, Postovit LM, Margaryan NV, Sam A, Hess AR, Wheaton WW et al. Embryonic and tumorigenic pathways converge via Nodal signaling: role in melanoma aggressiveness. Nat Med 2006; 12: 925–932. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

