Tensor decomposition for multiple-tissue gene expression experiments
- PMID: 27479908
- PMCID: PMC5010142
- DOI: 10.1038/ng.3624
Tensor decomposition for multiple-tissue gene expression experiments
Abstract
Genome-wide association studies of gene expression traits and other cellular phenotypes have successfully identified links between genetic variation and biological processes. The majority of discoveries have uncovered cis-expression quantitative trait locus (eQTL) effects via mass univariate testing of SNPs against gene expression in single tissues. Here we present a Bayesian method for multiple-tissue experiments focusing on uncovering gene networks linked to genetic variation. Our method decomposes the 3D array (or tensor) of gene expression measurements into a set of latent components. We identify sparse gene networks that can then be tested for association against genetic variation across the genome. We apply our method to a data set of 845 individuals from the TwinsUK cohort with gene expression measured via RNA-seq analysis in adipose, lymphoblastoid cell lines (LCLs) and skin. We uncover several gene networks with a genetic basis and clear biological and statistical significance. Extensions of this approach will allow integration of different omics, environmental and phenotypic data sets.
Figures


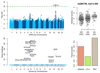




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

