Transcriptome Profiling of the Green Alga Spirogyra pratensis (Charophyta) Suggests an Ancestral Role for Ethylene in Cell Wall Metabolism, Photosynthesis, and Abiotic Stress Responses
- PMID: 27489312
- PMCID: PMC5074641
- DOI: 10.1104/pp.16.00299
Transcriptome Profiling of the Green Alga Spirogyra pratensis (Charophyta) Suggests an Ancestral Role for Ethylene in Cell Wall Metabolism, Photosynthesis, and Abiotic Stress Responses
Abstract
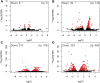
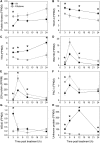
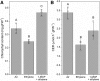
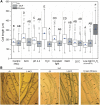
It is well known that ethylene regulates a diverse set of developmental and stress-related processes in angiosperms, yet its roles in early-diverging embryophytes and algae are poorly understood. Recently, it was shown that ethylene functions as a hormone in the charophyte green alga Spirogyra pratensis Since land plants evolved from charophytes, this implies conservation of ethylene as a hormone in green plants for at least 450 million years. However, the physiological role of ethylene in charophyte algae has remained unknown. To gain insight into ethylene responses in Spirogyra, we used mRNA sequencing to measure changes in gene expression over time in Spirogyra filaments in response to an ethylene treatment. Our analyses show that at the transcriptional level, ethylene predominantly regulates three processes in Spirogyra: (1) modification of the cell wall matrix by expansins and xyloglucan endotransglucosylases/hydrolases, (2) down-regulation of chlorophyll biosynthesis and photosynthesis, and (3) activation of abiotic stress responses. We confirmed that the photosynthetic capacity and chlorophyll content were reduced by an ethylene treatment and that several abiotic stress conditions could stimulate cell elongation in an ethylene-dependent manner. We also found that the Spirogyra transcriptome harbors only 10 ethylene-responsive transcription factor (ERF) homologs, several of which are regulated by ethylene. These results provide an initial understanding of the hormonal responses induced by ethylene in Spirogyra and help to reconstruct the role of ethylene in ancestral charophytes prior to the origin of land plants.
© 2016 American Society of Plant Biologists. All rights reserved.
Figures




Similar articles
-
Evidence for land plant cell wall biosynthetic mechanisms in charophyte green algae.Ann Bot. 2014 Oct;114(6):1217-36. doi: 10.1093/aob/mcu171. Epub 2014 Sep 9. Ann Bot. 2014. PMID: 25204387 Free PMC article.
-
Conservation of ethylene as a plant hormone over 450 million years of evolution.Nat Plants. 2015 Jan 8;1:14004. doi: 10.1038/nplants.2014.4. Nat Plants. 2015. PMID: 27246051
-
Search for evolutionary roots of land plant arabinogalactan-proteins in charophytes: presence of a rhamnogalactan-protein in Spirogyra pratensis (Zygnematophyceae).Plant J. 2022 Feb;109(3):568-584. doi: 10.1111/tpj.15577. Epub 2021 Nov 26. Plant J. 2022. PMID: 34767672 Free PMC article.
-
Xylans of Red and Green Algae: What Is Known about Their Structures and How They Are Synthesised?Polymers (Basel). 2019 Feb 18;11(2):354. doi: 10.3390/polym11020354. Polymers (Basel). 2019. PMID: 30960338 Free PMC article. Review.
-
Ethylene: A Master Regulator of Plant-Microbe Interactions under Abiotic Stresses.Cells. 2022 Dec 21;12(1):31. doi: 10.3390/cells12010031. Cells. 2022. PMID: 36611825 Free PMC article. Review.
Cited by
-
Induction of Conjugation and Zygospore Cell Wall Characteristics in the Alpine Spirogyra mirabilis (Zygnematophyceae, Charophyta): Advantage under Climate Change Scenarios?Plants (Basel). 2021 Aug 23;10(8):1740. doi: 10.3390/plants10081740. Plants (Basel). 2021. PMID: 34451785 Free PMC article.
-
On plant defense signaling networks and early land plant evolution.Commun Integr Biol. 2018 Aug 9;11(3):1-14. doi: 10.1080/19420889.2018.1486168. eCollection 2018. Commun Integr Biol. 2018. PMID: 30214675 Free PMC article. Review.
-
Cell Wall Enzymes in Zygnema circumcarinatum UTEX 1559 Respond to Osmotic Stress in a Plant-Like Fashion.Front Plant Sci. 2019 Jun 7;10:732. doi: 10.3389/fpls.2019.00732. eCollection 2019. Front Plant Sci. 2019. PMID: 31231410 Free PMC article.
-
Ethylene Exerts Species-Specific and Age-Dependent Control of Photosynthesis.Plant Physiol. 2018 Apr;176(4):2601-2612. doi: 10.1104/pp.17.01706. Epub 2018 Feb 2. Plant Physiol. 2018. PMID: 29438047 Free PMC article. Review.
-
Embryophyte stress signaling evolved in the algal progenitors of land plants.Proc Natl Acad Sci U S A. 2018 Apr 10;115(15):E3471-E3480. doi: 10.1073/pnas.1719230115. Epub 2018 Mar 26. Proc Natl Acad Sci U S A. 2018. PMID: 29581286 Free PMC article.
References
-
- Andersen RA, Berges JA, Harrison PJ, Watanabe MM (2005) Recipes for freshwater and saltwater media. In Andersen RA, Algal Culturing Techniques. Elsevier Academic Press, London, pp 429–538.
-
- Bashline L, Lei L, Li S, Gu Y (2014) Cell wall, cytoskeleton, and cell expansion in higher plants. Mol Plant 7: 586–600 - PubMed
-
- Bleecker AB, Estelle MA, Somerville C, Kende H (1988) Insensitivity to ethylene conferred by a dominant mutation in Arabidopsis thaliana. Science 241: 1086–1089 - PubMed
-
- Bremer K. (1985) Summary of green plants phylogeny and classification. Cladistics 1: 369–385 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

