Cleavage of Model Substrates by Arabidopsis thaliana PRORP1 Reveals New Insights into Its Substrate Requirements
- PMID: 27494328
- PMCID: PMC4975455
- DOI: 10.1371/journal.pone.0160246
Cleavage of Model Substrates by Arabidopsis thaliana PRORP1 Reveals New Insights into Its Substrate Requirements
Abstract
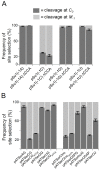
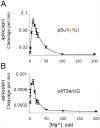
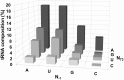
Two broad classes of RNase P trim the 5' leader of precursor tRNAs (pre-tRNAs): ribonucleoprotein (RNP)- and proteinaceous (PRORP)-variants. These two RNase P types, which use different scaffolds for catalysis, reflect independent evolutionary paths. While the catalytic RNA-based RNP form is present in all three domains of life, the PRORP family is restricted to eukaryotes. To obtain insights on substrate recognition by PRORPs, we examined the 5' processing ability of recombinant Arabidopsis thaliana PRORP1 (AtPRORP1) using a panel of pre-tRNASer variants and model hairpin-loop derivatives (pATSer type) that consist of the acceptor-T-stem stack and the T-/D-loop. Our data indicate the importance of the identity of N-1 (the residue immediately 5' to the cleavage site) and the N-1:N+73 base pair for cleavage rate and site selection of pre-tRNASer and pATSer. The nucleobase preferences that we observed mirror the frequency of occurrence in the complete suite of organellar pre-tRNAs in eight algae/plants that we analyzed. The importance of the T-/D-loop in pre-tRNASer for tight binding to AtPRORP1 is indicated by the 200-fold weaker binding of pATSer compared to pre-tRNASer, while the essentiality of the T-loop for cleavage is reflected by the near-complete loss of activity when a GAAA-tetraloop replaced the T-loop in pATSer. Substituting the 2'-OH at N-1 with 2'-H also resulted in no detectable cleavage, hinting at the possible role of this 2'-OH in coordinating Mg2+ ions critical for catalysis. Collectively, our results indicate similarities but also key differences in substrate recognition by the bacterial RNase P RNP and AtPRORP1: while both forms exploit the acceptor-T-stem stack and the elbow region in the pre-tRNA, the RNP form appears to require more recognition determinants for cleavage-site selection.
Conflict of interest statement
Figures
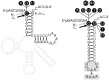


Similar articles
-
Molecular recognition of pre-tRNA by Arabidopsis protein-only Ribonuclease P.RNA. 2017 Dec;23(12):1860-1873. doi: 10.1261/rna.061457.117. Epub 2017 Sep 5. RNA. 2017. PMID: 28874505 Free PMC article.
-
Protein-only RNase P function in Escherichia coli: viability, processing defects and differences between PRORP isoenzymes.Nucleic Acids Res. 2017 Jul 7;45(12):7441-7454. doi: 10.1093/nar/gkx405. Nucleic Acids Res. 2017. PMID: 28499021 Free PMC article.
-
Pentatricopeptide repeat motifs in the processing enzyme PRORP1 in Arabidopsis thaliana play a crucial role in recognition of nucleotide bases at TψC loop in precursor tRNAs.Biochem Biophys Res Commun. 2014 Aug 8;450(4):1541-6. doi: 10.1016/j.bbrc.2014.07.030. Epub 2014 Jul 15. Biochem Biophys Res Commun. 2014. PMID: 25034328
-
PPR proteins shed a new light on RNase P biology.RNA Biol. 2013;10(9):1457-68. doi: 10.4161/rna.25273. Epub 2013 Jun 19. RNA Biol. 2013. PMID: 23925311 Free PMC article. Review.
-
RNase P RNA mediated cleavage: substrate recognition and catalysis.Biochimie. 2007 Oct;89(10):1183-94. doi: 10.1016/j.biochi.2007.05.009. Epub 2007 Jun 2. Biochimie. 2007. PMID: 17624654 Review.
Cited by
-
How RNases Shape Mitochondrial Transcriptomes.Int J Mol Sci. 2022 May 30;23(11):6141. doi: 10.3390/ijms23116141. Int J Mol Sci. 2022. PMID: 35682820 Free PMC article. Review.
-
RNase P: Beyond Precursor tRNA Processing.Genomics Proteomics Bioinformatics. 2024 May 9;22(1):qzae016. doi: 10.1093/gpbjnl/qzae016. Genomics Proteomics Bioinformatics. 2024. PMID: 38862431 Free PMC article. Review.
-
Importance of residue 248 in Escherichia coli RNase P RNA mediated cleavage.Sci Rep. 2023 Aug 29;13(1):14140. doi: 10.1038/s41598-023-41203-4. Sci Rep. 2023. PMID: 37644068 Free PMC article.
-
Synthetic riboswitches for the analysis of tRNA processing by eukaryotic RNase P enzymes.RNA. 2022 Apr;28(4):551-567. doi: 10.1261/rna.078814.121. Epub 2022 Jan 12. RNA. 2022. PMID: 35022261 Free PMC article.
-
Molecular recognition of pre-tRNA by Arabidopsis protein-only Ribonuclease P.RNA. 2017 Dec;23(12):1860-1873. doi: 10.1261/rna.061457.117. Epub 2017 Sep 5. RNA. 2017. PMID: 28874505 Free PMC article.
References
-
- Guerrier-Takada C., Gardiner K., Marsh T.,Pace N. and Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983;35, 849–857. - PubMed
-
- McClain W.H., Guerrier-Takada C. and Altman S. Model substrates for an RNA enzyme. Science. 1987;238, 527–530. - PubMed
-
- Forster A.C. and Altman S. External guide sequences for an RNA enzyme. Science. 1990;249, 783–786. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

