Mitochondrial Protein Interaction Mapping Identifies Regulators of Respiratory Chain Function
- PMID: 27499296
- PMCID: PMC4992456
- DOI: 10.1016/j.molcel.2016.06.033
Mitochondrial Protein Interaction Mapping Identifies Regulators of Respiratory Chain Function
Abstract
Mitochondria are essential for numerous cellular processes, yet hundreds of their proteins lack robust functional annotation. To reveal functions for these proteins (termed MXPs), we assessed condition-specific protein-protein interactions for 50 select MXPs using affinity enrichment mass spectrometry. Our data connect MXPs to diverse mitochondrial processes, including multiple aspects of respiratory chain function. Building upon these observations, we validated C17orf89 as a complex I (CI) assembly factor. Disruption of C17orf89 markedly reduced CI activity, and its depletion is found in an unresolved case of CI deficiency. We likewise discovered that LYRM5 interacts with and deflavinates the electron-transferring flavoprotein that shuttles electrons to coenzyme Q (CoQ). Finally, we identified a dynamic human CoQ biosynthetic complex involving multiple MXPs whose topology we map using purified components. Collectively, our data lend mechanistic insight into respiratory chain-related activities and prioritize hundreds of additional interactions for further exploration of mitochondrial protein function.
Keywords: C15orf48; C2orf47; DHRS4.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures



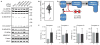

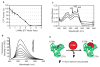

References
MeSH terms
Substances
Grants and funding
- F30 AG043282/AG/NIA NIH HHS/United States
- R01 GM112057/GM/NIGMS NIH HHS/United States
- T32 GM008692/GM/NIGMS NIH HHS/United States
- R35 GM118110/GM/NIGMS NIH HHS/United States
- NIHR-HCS-D12-03-04/DH_/Department of Health/United Kingdom
- G0601943/MRC_/Medical Research Council/United Kingdom
- R01 GM029076/GM/NIGMS NIH HHS/United States
- 096919/Z/11/Z/WT_/Wellcome Trust/United Kingdom
- T32 GM007215/GM/NIGMS NIH HHS/United States
- T15 LM007359/LM/NLM NIH HHS/United States
- R01 DK098672/DK/NIDDK NIH HHS/United States
- U01 GM094622/GM/NIGMS NIH HHS/United States
- R01 GM115591/GM/NIGMS NIH HHS/United States
- T32 HL007899/HL/NHLBI NIH HHS/United States
- WT_/Wellcome Trust/United Kingdom
- T32 DK007665/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

