Proteasome inhibition for treatment of leishmaniasis, Chagas disease and sleeping sickness
- PMID: 27501246
- PMCID: PMC5161665
- DOI: 10.1038/nature19339
Proteasome inhibition for treatment of leishmaniasis, Chagas disease and sleeping sickness
Abstract
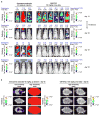
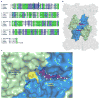
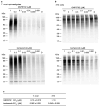
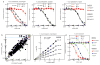
Chagas disease, leishmaniasis and sleeping sickness affect 20 million people worldwide and lead to more than 50,000 deaths annually. The diseases are caused by infection with the kinetoplastid parasites Trypanosoma cruzi, Leishmania spp. and Trypanosoma brucei spp., respectively. These parasites have similar biology and genomic sequence, suggesting that all three diseases could be cured with drugs that modulate the activity of a conserved parasite target. However, no such molecular targets or broad spectrum drugs have been identified to date. Here we describe a selective inhibitor of the kinetoplastid proteasome (GNF6702) with unprecedented in vivo efficacy, which cleared parasites from mice in all three models of infection. GNF6702 inhibits the kinetoplastid proteasome through a non-competitive mechanism, does not inhibit the mammalian proteasome or growth of mammalian cells, and is well-tolerated in mice. Our data provide genetic and chemical validation of the parasite proteasome as a promising therapeutic target for treatment of kinetoplastid infections, and underscore the possibility of developing a single class of drugs for these neglected diseases.
Conflict of interest statement
Patents related to this work has been filed (WO 2015/095477 A1, WO 2014/151784 A1, WO 2014/151729). Several authors own shares of Novartis.
Figures

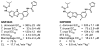


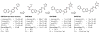

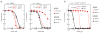
References
-
- World Health Organization. Research priorities for Chagas disease human African trypanosomiasis leishmaniasis. 2012. pp. 1–100. WHO Technical Report Series 975. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

