The Lysosome as a Regulatory Hub
- PMID: 27501449
- PMCID: PMC9345128
- DOI: 10.1146/annurev-cellbio-111315-125125
The Lysosome as a Regulatory Hub
Abstract
The lysosome has long been viewed as the recycling center of the cell. However, recent discoveries have challenged this simple view and have established a central role of the lysosome in nutrient-dependent signal transduction. The degradative role of the lysosome and its newly discovered signaling functions are not in conflict but rather cooperate extensively to mediate fundamental cellular activities such as nutrient sensing, metabolic adaptation, and quality control of proteins and organelles. Moreover, lysosome-based signaling and degradation are subject to reciprocal regulation. Transcriptional programs of increasing complexity control the biogenesis, composition, and abundance of lysosomes and fine-tune their activity to match the evolving needs of the cell. Alterations in these essential activities are, not surprisingly, central to the pathophysiology of an ever-expanding spectrum of conditions, including storage disorders, neurodegenerative diseases, and cancer. Thus, unraveling the functions of this fascinating organelle will contribute to our understanding of the fundamental logic of metabolic organization and will point to novel therapeutic avenues in several human diseases.
Keywords: TFEB; autophagy; cancer metabolism; lysosomal adaptation; mTORC1; neurodegeneration; nutrient sensing.
Figures


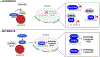
References
-
- Aharon-Peretz J, Rosenbaum H, Gershoni-Baruch R. 2004. Mutations in the glucocerebrosidase gene and Parkinson’s disease in Ashkenazi Jews. The New England journal of medicine 351: 1972–7 - PubMed
-
- Aits S, Jaattela M. 2013. Lysosomal cell death at a glance. Journal of cell science 126: 1905–12 - PubMed
-
- Ballabio A, Gieselmann V. 2009. Lysosomal disorders: from storage to cellular damage. Biochim Biophys Acta 1793: 684–96 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

