Spatial Genome Organization and Its Emerging Role as a Potential Diagnosis Tool
- PMID: 27507988
- PMCID: PMC4961005
- DOI: 10.3389/fgene.2016.00134
Spatial Genome Organization and Its Emerging Role as a Potential Diagnosis Tool
Abstract
In eukaryotic cells the genome is highly spatially organized. Functional relevance of higher order genome organization is implied by the fact that specific genes, and even whole chromosomes, alter spatial position in concert with functional changes within the nucleus, for example with modifications to chromatin or transcription. The exact molecular pathways that regulate spatial genome organization and the full implication to the cell of such an organization remain to be determined. However, there is a growing realization that the spatial organization of the genome can be used as a marker of disease. While global genome organization patterns remain largely conserved in disease, some genes and chromosomes occupy distinct nuclear positions in diseased cells compared to their normal counterparts, with the patterns of reorganization differing between diseases. Importantly, mapping the spatial positioning patterns of specific genomic loci can distinguish cancerous tissue from benign with high accuracy. Genome positioning is an attractive novel biomarker since additional quantitative biomarkers are urgently required in many cancer types. Current diagnostic techniques are often subjective and generally lack the ability to identify aggressive cancer from indolent, which can lead to over- or under-treatment of patients. Proof-of-principle for the use of genome positioning as a diagnostic tool has been provided based on small scale retrospective studies. Future large-scale studies are required to assess the feasibility of bringing spatial genome organization-based diagnostics to the clinical setting and to determine if the positioning patterns of specific loci can be useful biomarkers for cancer prognosis. Since spatial reorganization of the genome has been identified in multiple human diseases, it is likely that spatial genome positioning patterns as a diagnostic biomarker may be applied to many diseases.
Keywords: cancer; diagnosis; disease; gene positioning; genome organization; nuclear architecture; spatial positioning.
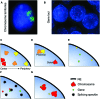
Figures

Similar articles
-
Assessment of the Utility of Gene Positioning Biomarkers in the Stratification of Prostate Cancers.Front Genet. 2019 Oct 17;10:1029. doi: 10.3389/fgene.2019.01029. eCollection 2019. Front Genet. 2019. PMID: 31681438 Free PMC article.
-
Causes and consequences of nuclear gene positioning.J Cell Sci. 2017 May 1;130(9):1501-1508. doi: 10.1242/jcs.199786. Epub 2017 Apr 12. J Cell Sci. 2017. PMID: 28404786 Free PMC article. Review.
-
Spatial genome organization.Exp Cell Res. 2004 May 15;296(1):64-70. doi: 10.1016/j.yexcr.2004.03.013. Exp Cell Res. 2004. PMID: 15120995 Review.
-
A predictive computational model of the dynamic 3D interphase yeast nucleus.Curr Biol. 2012 Oct 23;22(20):1881-90. doi: 10.1016/j.cub.2012.07.069. Epub 2012 Aug 30. Curr Biol. 2012. PMID: 22940469
-
Locus-specific gene repositioning in prostate cancer.Mol Biol Cell. 2016 Jan 15;27(2):236-46. doi: 10.1091/mbc.E15-05-0280. Epub 2015 Nov 12. Mol Biol Cell. 2016. PMID: 26564800 Free PMC article.
Cited by
-
Nuclear morphometrics and chromatin condensation patterns as disease biomarkers using a mobile microscope.PLoS One. 2019 Jul 17;14(7):e0218757. doi: 10.1371/journal.pone.0218757. eCollection 2019. PLoS One. 2019. PMID: 31314779 Free PMC article.
-
The snail Biomphalaria glabrata as a model to interrogate the molecular basis of complex human diseases.PLoS Negl Trop Dis. 2018 Aug 9;12(8):e0006552. doi: 10.1371/journal.pntd.0006552. eCollection 2018 Aug. PLoS Negl Trop Dis. 2018. PMID: 30091971 Free PMC article. No abstract available.
-
Assessment of the Utility of Gene Positioning Biomarkers in the Stratification of Prostate Cancers.Front Genet. 2019 Oct 17;10:1029. doi: 10.3389/fgene.2019.01029. eCollection 2019. Front Genet. 2019. PMID: 31681438 Free PMC article.
-
Large-scale mapping of positional changes of hypoxia-responsive genes upon activation.Mol Biol Cell. 2022 Jul 1;33(8):ar72. doi: 10.1091/mbc.E21-11-0593. Epub 2022 Apr 27. Mol Biol Cell. 2022. PMID: 35476603 Free PMC article.
-
Protein sequestration at the nuclear periphery as a potential regulatory mechanism in premature aging.J Cell Biol. 2018 Jan 2;217(1):21-37. doi: 10.1083/jcb.201706061. Epub 2017 Oct 19. J Cell Biol. 2018. PMID: 29051264 Free PMC article. Review.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

