Baculovirus FP25K Localization: Role of the Coiled-Coil Domain
- PMID: 27512078
- PMCID: PMC5068512
- DOI: 10.1128/JVI.01241-16
Baculovirus FP25K Localization: Role of the Coiled-Coil Domain
Abstract
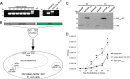
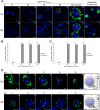
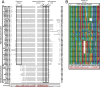
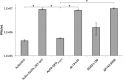
Two types of viruses are produced during the baculovirus life cycle: budded virus (BV) and occlusion-derived virus (ODV). A particular baculovirus protein, FP25K, is involved in the switch from BV to ODV production. Previously, FP25K from the model alphabaculovirus Autographa californica multiple nucleopolyhedrovirus (AcMNPV) was shown to traffic ODV envelope proteins. However, FP25K localization and the domains involved are inconclusive. Here we used a quantitative approach to study FP25K subcellular localization during infection using an AcMNPV bacmid virus that produces a functional AcMNPV FP25K-green fluorescent protein (GFP) fusion protein. During cell infection, FP25K-GFP localized primarily to the cytoplasm, particularly amorphous structures, with a small fraction being localized in the nucleus. To investigate the sequences involved in FP25K localization, an alignment of baculovirus FP25K sequences revealed that the N-terminal putative coiled-coil domain is present in all alphabaculoviruses but absent in betabaculoviruses. Structural prediction indicated a strong relatedness of AcMNPV FP25K to long interspersed element 1 (LINE-1) open reading frame 1 protein (ORF1p), which contains an N-terminal coiled-coil domain responsible for cytoplasmic retention. Point mutations and deletions of this domain lead to a change in AcMNPV FP25K localization from cytoplasmic to nuclear. The coiled-coil and C-terminal deletion viruses increased BV production. Furthermore, a betabaculovirus FP25K protein lacking this N-terminal coiled-coil domain localized predominantly to the nucleus and exhibited increased BV production. These data suggest that the acquisition of this N-terminal coiled-coil domain in FP25K is important for the evolution of alphabaculoviruses. Moreover, with the divergence of preocclusion nuclear membrane breakdown in betabaculoviruses and membrane integrity in alphabaculoviruses, this domain represents an alphabaculovirus adaptation for nuclear trafficking of occlusion-associated proteins.
Importance: Baculovirus infection produces two forms of viruses: BV and ODV. Manufacturing of ODV involves trafficking of envelope proteins to the inner nuclear membrane, mediated partly through the FP25K protein. Since FP25K is present in alpha-, beta-, and gammabaculoviruses, it is uncertain if this trafficking function is conserved. In this study, we looked at alpha- and betabaculovirus FP25K trafficking by its localization. Alphabaculovirus FP25K localized primarily to the cytoplasm, whereas betabaculovirus FP25K localized to the nucleus. We found that an N-terminal coiled-coil domain present in all alphabaculovirus FP25K proteins, but absent in betabaculovirus FP25K, was critical for alphabaculovirus FP25K cytoplasmic localization. We believe that this represents an evolutionary process that partly led to the gain of function of this N-terminal coiled-coil domain in alphabaculovirus FP25K to aid in nuclear trafficking of occlusion-associated proteins. Due to betabaculovirus breakdown of the nuclear membrane before occlusion, this function is not needed, and the domain was lost or never acquired.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures



Similar articles
-
Autographa californica nuclear polyhedrosis virus: subcellular localization and protein trafficking of BV/ODV-E26 to intranuclear membranes and viral envelopes.Virology. 1998 Jan 5;240(1):64-75. doi: 10.1006/viro.1997.8903. Virology. 1998. PMID: 9448690
-
Effects of deletion and overexpression of the Autographa californica nuclear polyhedrosis virus FP25K gene on synthesis of two occlusion-derived virus envelope proteins and their transport into virus-induced intranuclear membranes.J Virol. 2001 Nov;75(22):10829-42. doi: 10.1128/JVI.75.22.10829-10842.2001. J Virol. 2001. PMID: 11602724 Free PMC article.
-
AcMNPV ac143 (odv-e18) is essential for mediating budded virus production and is the 30th baculovirus core gene.Virology. 2008 May 25;375(1):277-91. doi: 10.1016/j.virol.2008.01.039. Epub 2008 Mar 6. Virology. 2008. PMID: 18328526
-
Molecular biology of the baculovirus occlusion-derived virus envelope.Curr Drug Targets. 2007 Oct;8(10):1084-95. doi: 10.2174/138945007782151315. Curr Drug Targets. 2007. PMID: 17979668 Review.
-
Baculovirus Entry and Egress from Insect Cells.Annu Rev Virol. 2018 Sep 29;5(1):113-139. doi: 10.1146/annurev-virology-092917-043356. Epub 2018 Jul 13. Annu Rev Virol. 2018. PMID: 30004832 Review.
Cited by
-
Improvement of protein production in baculovirus expression vector system by removing a total of 10 kb of nonessential fragments from Autographa californica multiple nucleopolyhedrovirus genome.Front Microbiol. 2023 Apr 13;14:1171500. doi: 10.3389/fmicb.2023.1171500. eCollection 2023. Front Microbiol. 2023. PMID: 37125202 Free PMC article.
-
A novel method for generating baculovirus bacmids using EGFP-mediated purification and linearization.Biotechnol Lett. 2025 Jul 17;47(4):77. doi: 10.1007/s10529-025-03619-y. Biotechnol Lett. 2025. PMID: 40670866
-
Identification and characterization of coiled-coil motifs across Autographa californica multiple nucleopolyhedrovirus genome.Heliyon. 2022 Sep 10;8(9):e10588. doi: 10.1016/j.heliyon.2022.e10588. eCollection 2022 Sep. Heliyon. 2022. PMID: 36132175 Free PMC article.
References
-
- Ikeda M, Hamajima R, Kobayashi M. 2015. Baculoviruses: diversity, evolution and manipulation of insects. Entomol Sci 18:1–20. doi:10.1111/ens.12105. - DOI
-
- Miller L. 1997. The baculoviruses. Plenum Press, New York, NY.
-
- Rohrmann GF. 2008. Baculovirus molecular biology. National Center for Biotechnology Information, Bethesda, MD. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

