Orthologs, turn-over, and remolding of tRNAs in primates and fruit flies
- PMID: 27515907
- PMCID: PMC4981973
- DOI: 10.1186/s12864-016-2927-4
Orthologs, turn-over, and remolding of tRNAs in primates and fruit flies
Abstract
Background: Transfer RNAs (tRNAs) are ubiquitous in all living organism. They implement the genetic code so that most genomes contain distinct tRNAs for almost all 61 codons. They behave similar to mobile elements and proliferate in genomes spawning both local and non-local copies. Most tRNA families are therefore typically present as multicopy genes. The members of the individual tRNA families evolve under concerted or rapid birth-death evolution, so that paralogous copies maintain almost identical sequences over long evolutionary time-scales. To a good approximation these are functionally equivalent. Individual tRNA copies thus are evolutionary unstable and easily turn into pseudogenes and disappear. This leads to a rapid turnover of tRNAs and often large differences in the tRNA complements of closely related species. Since tRNA paralogs are not distinguished by sequence, common methods cannot not be used to establish orthology between tRNA genes.
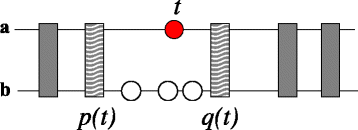
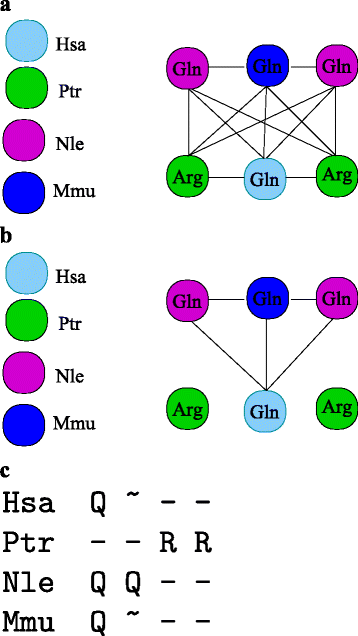
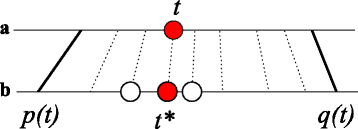
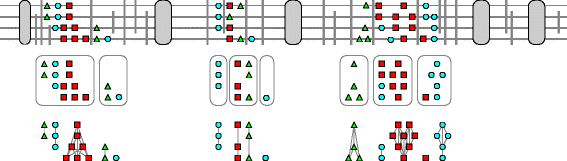
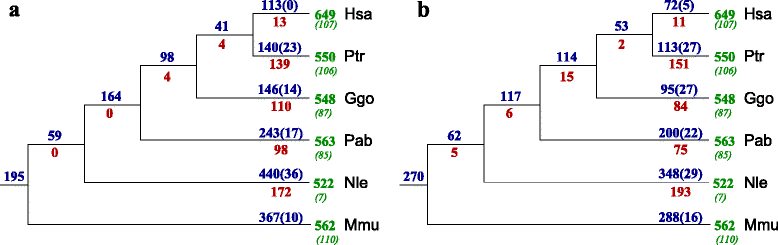
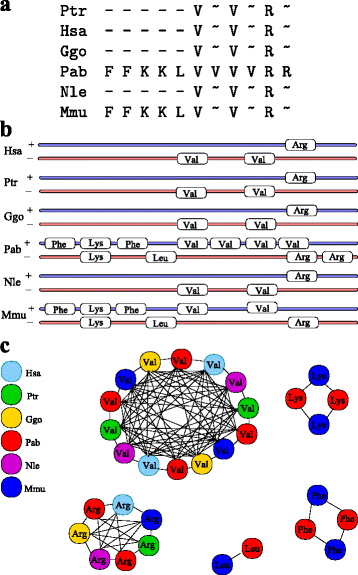
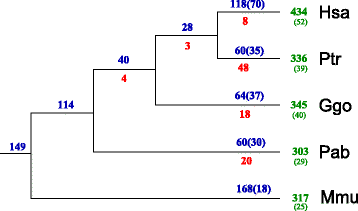
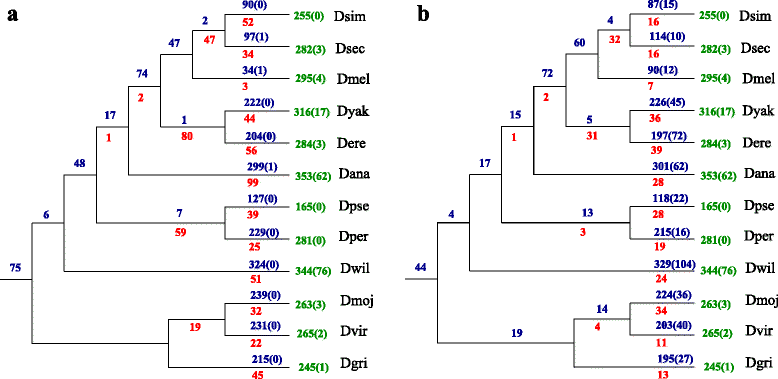
Results: In this contribution we introduce a general framework to distinguish orthologs and paralogs in gene families that are subject to concerted evolution. It is based on the use of uniquely aligned adjacent sequence elements as anchors to establish syntenic conservation of sequence intervals. In practice, anchors and intervals can be extracted from genome-wide multiple sequence alignments. Syntenic clusters of concertedly evolving genes of different families can then be subdivided by list alignments, leading to usually small clusters of candidate co-orthologs. On the basis of recent advances in phylogenetic combinatorics, these candidate clusters can be further processed by cograph editing to recover their duplication histories. We developed a workflow that can be conceptualized as stepwise refinement of a graph of homologous genes. We apply this analysis strategy with different types of synteny anchors to investigate the evolution of tRNAs in primates and fruit flies. We identified a large number of tRNA remolding events concentrated at the tips of the phylogeny. With one notable exception all phylogenetically old tRNA remoldings do not change the isoacceptor class.
Conclusions: Gene families evolving under concerted evolution are not amenable to classical phylogenetic analyses since paralogs maintain identical, species-specific sequences, precluding the estimation of correct gene trees from sequence differences. This leaves conservation of syntenic arrangements with respect to "anchor elements" that are not subject to concerted evolution as the only viable source of phylogenetic information. We have demonstrated here that a purely synteny-based analysis of tRNA gene histories is indeed feasible. Although the choice of synteny anchors influences the resolution in particular when tight gene clusters are present, and the quality of sequence alignments, genome assemblies, and genome rearrangements limits the scope of the analysis, largely coherent results can be obtained for tRNAs. In particular, we conclude that a large fraction of the tRNAs are recent copies. This proliferation is compensated by rapid pseudogenization as exemplified by many very recent alloacceptor remoldings.
Keywords: Concerted evolution; Orthology; Synteny; tRNA remolding.
Figures

Similar articles
-
Towards a comprehensive picture of alloacceptor tRNA remolding in metazoan mitochondrial genomes.Nucleic Acids Res. 2015 Sep 18;43(16):8044-56. doi: 10.1093/nar/gkv746. Epub 2015 Jul 30. Nucleic Acids Res. 2015. PMID: 26227972 Free PMC article.
-
Genomic organization of eukaryotic tRNAs.BMC Genomics. 2010 Apr 28;11:270. doi: 10.1186/1471-2164-11-270. BMC Genomics. 2010. PMID: 20426822 Free PMC article.
-
SMORE: Synteny Modulator of Repetitive Elements.Life (Basel). 2017 Oct 31;7(4):42. doi: 10.3390/life7040042. Life (Basel). 2017. PMID: 29088079 Free PMC article.
-
The evolving tRNA molecule.CRC Crit Rev Biochem. 1981;11(1):35-104. doi: 10.3109/10409238109108699. CRC Crit Rev Biochem. 1981. PMID: 7030617 Review.
-
Interchangeable parts: The evolutionarily dynamic tRNA population in plant mitochondria.Mitochondrion. 2020 May;52:144-156. doi: 10.1016/j.mito.2020.03.007. Epub 2020 Mar 14. Mitochondrion. 2020. PMID: 32184120 Review.
Cited by
-
Unbiased anchors for reliable genome-wide synteny detection.Algorithms Mol Biol. 2025 Apr 5;20(1):5. doi: 10.1186/s13015-025-00275-9. Algorithms Mol Biol. 2025. PMID: 40188341 Free PMC article.
-
Toward a Categorization of Virus-ncRNA Interactions in the World of RNA to Disentangle the Tiny Secrets of Dengue Virus.Viruses. 2024 May 18;16(5):804. doi: 10.3390/v16050804. Viruses. 2024. PMID: 38793685 Free PMC article. Review.
-
Evolution of genome structure in the Drosophila simulans species complex.Genome Res. 2021 Mar;31(3):380-396. doi: 10.1101/gr.263442.120. Epub 2021 Feb 9. Genome Res. 2021. PMID: 33563718 Free PMC article.
-
A tRNA gene potential to activate interferon signaling involves selective termination and is suppressible by La protein/SSB.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf513. doi: 10.1093/nar/gkaf513. Nucleic Acids Res. 2025. PMID: 40637226 Free PMC article.
-
miRNAture-Computational Detection of microRNA Candidates.Genes (Basel). 2021 Feb 27;12(3):348. doi: 10.3390/genes12030348. Genes (Basel). 2021. PMID: 33673400 Free PMC article.
References
-
- Fitch WM. Distinguishing homologous from analogous proteins. Syst Biol. 1970;19:99–113. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

