Effect of internal cleavage site mutations in human immunodeficiency virus type 1 capsid protein on its structure and function
- PMID: 27516963
- PMCID: PMC4971840
- DOI: 10.1002/2211-5463.12094
Effect of internal cleavage site mutations in human immunodeficiency virus type 1 capsid protein on its structure and function
Abstract
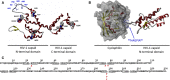
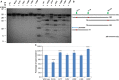
The capsid protein of the human immunodeficiency virus type 1 has been found to be a substrate of the retroviral protease in vitro, and its processing was predicted to be strongly dependent on a pH-induced conformational change. Several protease cleavage sites have been identified within the capsid protein, but the importance of its cleavage by the viral protease at the early phase of infection is controversial. To confirm the relevance of this process, we aimed to design, produce, and characterize mutant capsid proteins, in which the protein susceptibility toward HIV-1 protease is altered without affecting other steps of the viral life cycle. Our results indicate that while the introduced mutations changed the cleavage rate at the mutated sites of the capsid protein by HIV-1 protease, some of them caused only negligible or moderate structural changes (A78V, L189F, and L189I). However, the effects of other mutations (W23A, A77P, and L189P) were dramatic, as assessed by secondary structure determination or cyclophilin A-binding assay. Based on our observations, the L189F mutant capsid remains structurally and functionally unchanged and may therefore be the best candidate for use in studies aimed at better understanding the role of the protease in the early postentry events of viral infection or retrovirus-mediated gene transduction.
Keywords: HIV‐1; capsid protein; circular dichroism spectroscopy; cyclophilin A; human immunodeficiency virus type 1; protease, mutagenesis.
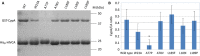
Figures

Similar articles
-
Human immunodeficiency virus type 1 capsid protein is a substrate of the retroviral proteinase while integrase is resistant toward proteolysis.Virology. 2003 May 25;310(1):16-23. doi: 10.1016/s0042-6822(03)00074-6. Virology. 2003. PMID: 12788626
-
Role of capsid sequence and immature nucleocapsid proteins p9 and p15 in Human Immunodeficiency Virus type 1 genomic RNA dimerization.Virology. 2009 Mar 1;385(1):233-44. doi: 10.1016/j.virol.2008.11.028. Epub 2008 Dec 13. Virology. 2009. PMID: 19070880
-
Naturally occurring amino acid polymorphisms in human immunodeficiency virus type 1 (HIV-1) Gag p7(NC) and the C-cleavage site impact Gag-Pol processing by HIV-1 protease.Virology. 2002 Jan 5;292(1):137-49. doi: 10.1006/viro.2001.1184. Virology. 2002. PMID: 11878916
-
Context-Dependent Cleavage of the Capsid Protein by the West Nile Virus Protease Modulates the Efficiency of Virus Assembly.J Virol. 2015 Aug;89(16):8632-42. doi: 10.1128/JVI.01253-15. Epub 2015 Jun 10. J Virol. 2015. PMID: 26063422 Free PMC article.
-
Natural variation in HIV-1 protease, Gag p7 and p6, and protease cleavage sites within gag/pol polyproteins: amino acid substitutions in the absence of protease inhibitors in mothers and children infected by human immunodeficiency virus type 1.Virology. 1996 May 15;219(2):407-16. doi: 10.1006/viro.1996.0266. Virology. 1996. PMID: 8638406
Cited by
-
Different Mutation Tolerance of Lentiviral (HIV-1) and Deltaretroviral (BLV and HTLV) Protease Precursors.Viruses. 2022 Aug 26;14(9):1888. doi: 10.3390/v14091888. Viruses. 2022. PMID: 36146695 Free PMC article.
References
-
- Tang S, Ablan S, Dueck M, Ayala‐López W, Soto B, Caplan M, Nagashima K, Hewlett IK, Freed EO and Levin JG (2006) A second‐site suppressor significantly improves the defective phenotype imposed by mutation of an aromatic residue in the N‐terminal domain of the HIV‐1 capsid protein. Virology 359, 105–115. - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

