PARP3 is a sensor of nicked nucleosomes and monoribosylates histone H2B(Glu2)
- PMID: 27530147
- PMCID: PMC4992063
- DOI: 10.1038/ncomms12404
PARP3 is a sensor of nicked nucleosomes and monoribosylates histone H2B(Glu2)
Abstract
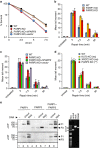
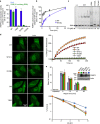
PARP3 is a member of the ADP-ribosyl transferase superfamily that we show accelerates the repair of chromosomal DNA single-strand breaks in avian DT40 cells. Two-dimensional nuclear magnetic resonance experiments reveal that PARP3 employs a conserved DNA-binding interface to detect and stably bind DNA breaks and to accumulate at sites of chromosome damage. PARP3 preferentially binds to and is activated by mononucleosomes containing nicked DNA and which target PARP3 trans-ribosylation activity to a single-histone substrate. Although nicks in naked DNA stimulate PARP3 autoribosylation, nicks in mononucleosomes promote the trans-ribosylation of histone H2B specifically at Glu2. These data identify PARP3 as a molecular sensor of nicked nucleosomes and demonstrate, for the first time, the ribosylation of chromatin at a site-specific DNA single-strand break.
Figures



Comment in
-
Nick Your DNA, Mark Your Chromatin.Mol Cell. 2016 Oct 6;64(1):7-9. doi: 10.1016/j.molcel.2016.09.023. Mol Cell. 2016. PMID: 27716488
Similar articles
-
Characterization of DNA ADP-ribosyltransferase activities of PARP2 and PARP3: new insights into DNA ADP-ribosylation.Nucleic Acids Res. 2018 Mar 16;46(5):2417-2431. doi: 10.1093/nar/gkx1318. Nucleic Acids Res. 2018. PMID: 29361132 Free PMC article.
-
Nick Your DNA, Mark Your Chromatin.Mol Cell. 2016 Oct 6;64(1):7-9. doi: 10.1016/j.molcel.2016.09.023. Mol Cell. 2016. PMID: 27716488
-
Sequential ADP-ribosylation pattern of nucleosomal histones. ADP-ribosylation of nucleosomal histones.Eur J Biochem. 1985 Jan 15;146(2):277-85. doi: 10.1111/j.1432-1033.1985.tb08650.x. Eur J Biochem. 1985. PMID: 2981686
-
PARP Power: A Structural Perspective on PARP1, PARP2, and PARP3 in DNA Damage Repair and Nucleosome Remodelling.Int J Mol Sci. 2021 May 12;22(10):5112. doi: 10.3390/ijms22105112. Int J Mol Sci. 2021. PMID: 34066057 Free PMC article. Review.
-
Relation between carcinogenesis, chromatin structure and poly(ADP-ribosylation) (review).Anticancer Res. 1991 Mar-Apr;11(2):489-527. Anticancer Res. 1991. PMID: 1905900 Review.
Cited by
-
Meta-analysis of DNA double-strand break response kinetics.Nucleic Acids Res. 2017 Dec 15;45(22):12625-12637. doi: 10.1093/nar/gkx1128. Nucleic Acids Res. 2017. PMID: 29182755 Free PMC article.
-
Structural basis of gap-filling DNA synthesis in the nucleosome by DNA Polymerase β.Nat Commun. 2025 Mar 17;16(1):2607. doi: 10.1038/s41467-025-57915-2. Nat Commun. 2025. PMID: 40097433 Free PMC article.
-
ADP-ribosylation: from molecular mechanisms to human disease.Genet Mol Biol. 2019 Dec 13;43(1 suppl 1):e20190075. doi: 10.1590/1678-4685-GMB-2019-0075. eCollection 2019. Genet Mol Biol. 2019. PMID: 31930280 Free PMC article.
-
Characterization of DNA ADP-ribosyltransferase activities of PARP2 and PARP3: new insights into DNA ADP-ribosylation.Nucleic Acids Res. 2018 Mar 16;46(5):2417-2431. doi: 10.1093/nar/gkx1318. Nucleic Acids Res. 2018. PMID: 29361132 Free PMC article.
-
PARP3 Affects Nucleosome Compaction Regulation.Int J Mol Sci. 2023 May 20;24(10):9042. doi: 10.3390/ijms24109042. Int J Mol Sci. 2023. PMID: 37240388 Free PMC article.
References
-
- Caldecott K. W. Single-strand break repair and genetic disease. Nat. Rev. Genet. 9, 619–631 (2008). - PubMed
-
- Caldecott K. W. XRCC1 and DNA strand break repair. DNA Repair (Amst.) 2, 955–969 (2003). - PubMed
-
- Rulten S. L. & Caldecott K. W. DNA strand break repair and neurodegeneration. DNA Repair (Amst.) 12, 558–567 (2013). - PubMed
-
- Caldecott K. W. Protein ADP-ribosylation and the cellular response to DNA strand breaks. DNA Repair (Amst.) 19, 108–113 (2014). - PubMed
-
- Hottiger M. O., Hassa P. O., Lüscher B., Schüler H. & Koch-Nolte F. Toward a unified nomenclature for mammalian ADP-ribosyltransferases. Trends Biochem. Sci. 35, 208–219 (2010). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

