Pyrophosphate-Dependent ATP Formation from Acetyl Coenzyme A in Syntrophus aciditrophicus, a New Twist on ATP Formation
- PMID: 27531911
- PMCID: PMC4992975
- DOI: 10.1128/mBio.01208-16
Pyrophosphate-Dependent ATP Formation from Acetyl Coenzyme A in Syntrophus aciditrophicus, a New Twist on ATP Formation
Abstract
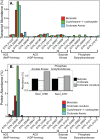
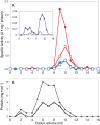
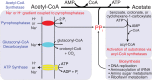
Syntrophus aciditrophicus is a model syntrophic bacterium that degrades key intermediates in anaerobic decomposition, such as benzoate, cyclohexane-1-carboxylate, and certain fatty acids, to acetate when grown with hydrogen-/formate-consuming microorganisms. ATP formation coupled to acetate production is the main source for energy conservation by S. aciditrophicus However, the absence of homologs for phosphate acetyltransferase and acetate kinase in the genome of S. aciditrophicus leaves it unclear as to how ATP is formed, as most fermentative bacteria rely on these two enzymes to synthesize ATP from acetyl coenzyme A (CoA) and phosphate. Here, we combine transcriptomic, proteomic, metabolite, and enzymatic approaches to show that S. aciditrophicus uses AMP-forming, acetyl-CoA synthetase (Acs1) for ATP synthesis from acetyl-CoA. acs1 mRNA and Acs1 were abundant in transcriptomes and proteomes, respectively, of S. aciditrophicus grown in pure culture and coculture. Cell extracts of S. aciditrophicus had low or undetectable acetate kinase and phosphate acetyltransferase activities but had high acetyl-CoA synthetase activity under all growth conditions tested. Both Acs1 purified from S. aciditrophicus and recombinantly produced Acs1 catalyzed ATP and acetate formation from acetyl-CoA, AMP, and pyrophosphate. High pyrophosphate levels and a high AMP-to-ATP ratio (5.9 ± 1.4) in S. aciditrophicus cells support the operation of Acs1 in the acetate-forming direction. Thus, S. aciditrophicus has a unique approach to conserve energy involving pyrophosphate, AMP, acetyl-CoA, and an AMP-forming, acetyl-CoA synthetase.
Importance: Bacteria use two enzymes, phosphate acetyltransferase and acetate kinase, to make ATP from acetyl-CoA, while acetate-forming archaea use a single enzyme, an ADP-forming, acetyl-CoA synthetase, to synthesize ATP and acetate from acetyl-CoA. Syntrophus aciditrophicus apparently relies on a different approach to conserve energy during acetyl-CoA metabolism, as its genome does not have homologs to the genes for phosphate acetyltransferase and acetate kinase. Here, we show that S. aciditrophicus uses an alternative approach, an AMP-forming, acetyl-CoA synthetase, to make ATP from acetyl-CoA. AMP-forming, acetyl-CoA synthetases were previously thought to function only in the activation of acetate to acetyl-CoA.
Copyright © 2016 James et al.
Figures
Similar articles
-
Syntrophus aciditrophicus uses the same enzymes in a reversible manner to degrade and synthesize aromatic and alicyclic acids.Environ Microbiol. 2019 May;21(5):1833-1846. doi: 10.1111/1462-2920.14601. Environ Microbiol. 2019. PMID: 30895699 Free PMC article.
-
Metabolism of benzoate, cyclohex-1-ene carboxylate, and cyclohexane carboxylate by "Syntrophus aciditrophicus" strain SB in syntrophic association with H(2)-using microorganisms.Appl Environ Microbiol. 2001 Apr;67(4):1728-38. doi: 10.1128/AEM.67.4.1728-1738.2001. Appl Environ Microbiol. 2001. PMID: 11282627 Free PMC article.
-
Benzoate fermentation by the anaerobic bacterium Syntrophus aciditrophicus in the absence of hydrogen-using microorganisms.Appl Environ Microbiol. 2001 Dec;67(12):5520-5. doi: 10.1128/AEM.67.12.5520-5525.2001. Appl Environ Microbiol. 2001. PMID: 11722901 Free PMC article.
-
Fermentative Cyclohexane Carboxylate Formation in Syntrophus aciditrophicus.J Mol Microbiol Biotechnol. 2016;26(1-3):165-79. doi: 10.1159/000440881. Epub 2016 Mar 10. J Mol Microbiol Biotechnol. 2016. PMID: 26959729 Review.
-
Acetate/acetyl-CoA metabolism associated with cancer fatty acid synthesis: overview and application.Cancer Lett. 2015 Jan 28;356(2 Pt A):211-6. doi: 10.1016/j.canlet.2014.02.019. Epub 2014 Feb 22. Cancer Lett. 2015. PMID: 24569091 Review.
Cited by
-
Comparison of muscle structure and transcriptome analysis of eyed-side muscle and blind-side muscle in Cynoglossussemilaevis (Osteichthyes, Cynoglossidae).Zookeys. 2025 Mar 6;1230:213-229. doi: 10.3897/zookeys.1230.139837. eCollection 2025. Zookeys. 2025. PMID: 40093692 Free PMC article. Review.
-
A New Pathway for Forming Acetate and Synthesizing ATP during Fermentation in Bacteria.Appl Environ Microbiol. 2021 Jun 25;87(14):e0295920. doi: 10.1128/AEM.02959-20. Epub 2021 Jun 25. Appl Environ Microbiol. 2021. PMID: 33931420 Free PMC article.
-
Synergy and competition during the anaerobic degradation of N-acetylglucosamine in a methane-emitting, subarctic, pH-neutral fen.Front Microbiol. 2024 Dec 11;15:1428517. doi: 10.3389/fmicb.2024.1428517. eCollection 2024. Front Microbiol. 2024. PMID: 39726964 Free PMC article.
-
Enoyl-Coenzyme A Respiration via Formate Cycling in Syntrophic Bacteria.mBio. 2021 Feb 22;13(1):e0374021. doi: 10.1128/mbio.03740-21. Epub 2022 Feb 1. mBio. 2021. PMID: 35100874 Free PMC article.
-
Concerted Metabolic Shifts Give New Insights Into the Syntrophic Mechanism Between Propionate-Fermenting Pelotomaculum thermopropionicum and Hydrogenotrophic Methanocella conradii.Front Microbiol. 2018 Jul 9;9:1551. doi: 10.3389/fmicb.2018.01551. eCollection 2018. Front Microbiol. 2018. PMID: 30038609 Free PMC article.
References
-
- Elshahed MS, Bhupathiraju VK, Wofford NQ, Nanny MA, McInerney MJ. 2001. Metabolism of benzoate, cyclohex-1-ene carboxylate, and cyclohexane carboxylate by Syntrophus aciditrophicus strain SB in syntrophic association with hydrogen-using microorganisms. Appl Environ Microbiol 67:1728–1738. doi:10.1128/AEM.67.4.1728-1738.2001. - DOI - PMC - PubMed
-
- McInerney MJ, Rohlin L, Mouttaki H, Kim U, Krupp RS, Rios-Hernandez L, Sieber J, Struchtemeyer CG, Bhattacharyya A, Campbell JW, Gunsalus RP. 2007. The genome of Syntrophus aciditrophicus: life at the thermodynamic limit of microbial growth. Proc Natl Acad Sci U S A 104:7600–7605. doi:10.1073/pnas.0610456104. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

