Blood-Borne Lipopolysaccharide Is Rapidly Eliminated by Liver Sinusoidal Endothelial Cells via High-Density Lipoprotein
- PMID: 27534554
- PMCID: PMC5010928
- DOI: 10.4049/jimmunol.1600702
Blood-Borne Lipopolysaccharide Is Rapidly Eliminated by Liver Sinusoidal Endothelial Cells via High-Density Lipoprotein
Abstract
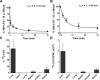
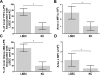
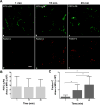
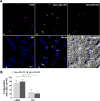
During Gram-negative bacterial infections, excessive LPS induces inflammation and sepsis via action on immune cells. However, the bulk of LPS can be cleared from circulation by the liver. Liver clearance is thought to be a slow process mediated exclusively by phagocytic resident macrophages, Kupffer cells (KC). However, we discovered that LPS disappears rapidly from the circulation, with a half-life of 2-4 min in mice, and liver eliminates about three quarters of LPS from blood circulation. Using microscopic techniques, we found that ∼75% of fluor-tagged LPS in liver became associated with liver sinusoidal endothelial cells (LSEC) and only ∼25% with KC. Notably, the ratio of LSEC-KC-associated LPS remained unchanged 45 min after infusion, indicating that LSEC independently processes the LPS. Most interestingly, results of kinetic analysis of LPS bioactivity, using modified limulus amebocyte lysate assay, suggest that recombinant factor C, an LPS binding protein, competitively inhibits high-density lipoprotein (HDL)-mediated LPS association with LSEC early in the process. Supporting the previous notion, 3 min postinfusion, 75% of infused fluorescently tagged LPS-HDL complex associates with LSEC, suggesting that HDL facilitates LPS clearance. These results lead us to propose a new paradigm of LSEC and HDL in clearing LPS with a potential to avoid inflammation during sepsis.
Copyright © 2016 by The American Association of Immunologists, Inc.
Figures




References
-
- Park BS, Song DH, Kim HM, Choi B-S, Lee H, Lee J-O. The structural basis of lipopolysaccharide recognition by the TLR4–MD-2 complex. Nature. 2009;458:1191–1196. - PubMed
-
- Freudenberg MA, Galanos C. Bacterial lipopolysaccharides: Structure, metabolism and mechanisms of action. Intern Rev Immunol. 1990;6:207–221. - PubMed
-
- Mathison JC, Ulevitch RJ. The Clearance, Tissue Distribution, and Cellular Localization of Intravenously Injected Lipopolysaccharide in Rabbits. J Immunol. 1979;123:2133–2143. - PubMed
-
- Shao B, Lu M, Katz SC, Varley AW, Hardwick J, Rogers TE, Ojogun N, Rockey DC, Dematteo RP, Munford RS. A host lipase detoxifies bacterial lipopolysaccharides in the liver and spleen. J Biol Chem. 2007;282:13726–13735. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

