Bolaamphiphile-based nanocomplex delivery of phosphorothioate gapmer antisense oligonucleotides as a treatment for Clostridium difficile
- PMID: 27536102
- PMCID: PMC4975145
- DOI: 10.2147/IJN.S109600
Bolaamphiphile-based nanocomplex delivery of phosphorothioate gapmer antisense oligonucleotides as a treatment for Clostridium difficile
Abstract
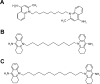
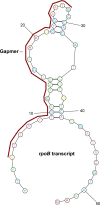
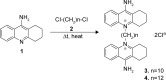
Despite being a conceptually appealing alternative to conventional antibiotics, a major challenge toward the successful implementation of antisense treatments for bacterial infections is the development of efficient oligonucleotide delivery systems. Cationic vesicles (bolasomes) composed of dequalinium chloride ("DQAsomes") have been used to deliver plasmid DNA across the cardiolipin-rich inner membrane of mitochondria. As cardiolipin is also a component of many bacterial membranes, we investigated the application of cationic bolasomes to bacteria as an oligonucleotide delivery system. Antisense sequences designed in silico to target the expression of essential genes of the bacterial pathogen, Clostridium difficile, were synthesized as 2'-O-methyl phosphorothioate gapmer antisense oligonucleotides (ASO). These antisense gapmers were quantitatively assessed for their ability to block mRNA translation using luciferase reporter and C. difficile protein expression plasmid constructs in a coupled transcription-translation system. Cationic bolaamphiphile compounds (dequalinium derivatives) of varying alkyl chain length were synthesized and bolasomes were prepared via probe sonication of an aqueous suspension. Bolasomes were characterized by particle size distribution, zeta potential, and binding capacities for anionic oligonucleotide. Bolasomes and antisense gapmers were combined to form antisense nanocomplexes. Anaerobic C. difficile log phase cultures were treated with serial doses of gapmer nanocomplexes or equivalent amounts of empty bolasomes for 24 hours. Antisense gapmers for four gene targets achieved nanomolar minimum inhibitory concentrations for C. difficile, with the lowest values observed for oligonucleotides targeting polymerase genes rpoB and dnaE. No inhibition of bacterial growth was observed from treatments at matched dosages of scrambled gapmer nanocomplexes or plain, oligonucleotide-free bolasomes compared to untreated control cultures. We describe the novel application of cationic bolasomes to deliver ASOs into bacteria. We also report the first successful in vitro antisense treatment to inhibit the growth of C. difficile.
Keywords: Clostridium difficile; antisense; bacteria; cationic bolaamphiphiles; dequalinium derivatives; gapmers; nanocomplex.
Figures






Similar articles
-
An Investigation into the Potential of Targeting Escherichia coli rne mRNA with Locked Nucleic Acid (LNA) Gapmers as an Antibacterial Strategy.Molecules. 2021 Jun 4;26(11):3414. doi: 10.3390/molecules26113414. Molecules. 2021. PMID: 34200016 Free PMC article.
-
Cationic amphiphilic bolaamphiphile-based delivery of antisense oligonucleotides provides a potentially microbiome sparing treatment for C. difficile.J Antibiot (Tokyo). 2018 Aug;71(8):713-721. doi: 10.1038/s41429-018-0056-9. Epub 2018 Apr 19. J Antibiot (Tokyo). 2018. PMID: 29674636 Free PMC article.
-
RNA Reduction and Hepatotoxic Potential Caused by Non-Gapmer Antisense Oligonucleotides.Nucleic Acid Ther. 2019 Feb;29(1):44-50. doi: 10.1089/nat.2018.0741. Epub 2018 Dec 1. Nucleic Acid Ther. 2019. PMID: 30508397
-
Oligonucleotide treatment of ras-induced tumors in nude mice.Mol Biotechnol. 2001 May;18(1):35-55. doi: 10.1385/MB:18:1:35. Mol Biotechnol. 2001. PMID: 11439698 Review.
-
[Mechanism for suppression mRNA translation with antisense oligonucleotides].Mol Biol (Mosk). 1990 Sep-Oct;24(5):1157-61. Mol Biol (Mosk). 1990. PMID: 2290415 Review. Russian.
Cited by
-
A comparative analysis of peptide-delivered antisense antibiotics using diverse nucleotide mimics.RNA. 2024 May 16;30(6):624-643. doi: 10.1261/rna.079969.124. RNA. 2024. PMID: 38413166 Free PMC article.
-
An Investigation into the Potential of Targeting Escherichia coli rne mRNA with Locked Nucleic Acid (LNA) Gapmers as an Antibacterial Strategy.Molecules. 2021 Jun 4;26(11):3414. doi: 10.3390/molecules26113414. Molecules. 2021. PMID: 34200016 Free PMC article.
-
Biomedically Relevant Applications of Bolaamphiphiles and Bolaamphiphile-Containing Materials.Front Chem. 2021 Jan 20;8:604151. doi: 10.3389/fchem.2020.604151. eCollection 2020. Front Chem. 2021. PMID: 33553103 Free PMC article. Review.
-
Promising strategies employing nucleic acids as antimicrobial drugs.Mol Ther Nucleic Acids. 2024 Jan 18;35(1):102122. doi: 10.1016/j.omtn.2024.102122. eCollection 2024 Mar 12. Mol Ther Nucleic Acids. 2024. PMID: 38333674 Free PMC article. Review.
-
Anti-virulence strategies for Clostridioides difficile infection: advances and roadblocks.Gut Microbes. 2020 Nov 9;12(1):1802865. doi: 10.1080/19490976.2020.1802865. Gut Microbes. 2020. PMID: 33092487 Free PMC article. Review.
References
-
- Davies AH, Roberts AK, Shone CC, Acharya KR. Super toxins from a super bug: structure and function of Clostridium difficile toxins. Biochem J. 2011;436(3):517–526. - PubMed
-
- McFarland LV, Elmer GW, Surawicz CM. Breaking the cycle: treatment strategies for 163 cases of recurrent Clostridium difficile disease. Am J Gastroenterol. 2002;97(7):1769–1775. - PubMed
-
- Cammarota G, Ianiro G, Gasbarrini A. Fecal microbiota transplantation for the treatment of Clostridium difficile infection: a systematic review. J Clin Gastroenterol. 2014;48(8):693–702. - PubMed
-
- Gilliland C, Weaver J, Stevenson M, et al. Association of Clostridium difficile ribotype 078 with detectable toxin in human stool specimens. J Med Microbiol. 2015;64(11):1341–1345. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

