Control of Anther Cell Differentiation by the Small Protein Ligand TPD1 and Its Receptor EMS1 in Arabidopsis
- PMID: 27537183
- PMCID: PMC4990239
- DOI: 10.1371/journal.pgen.1006147
Control of Anther Cell Differentiation by the Small Protein Ligand TPD1 and Its Receptor EMS1 in Arabidopsis
Abstract
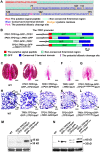
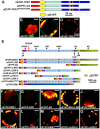
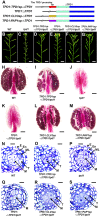
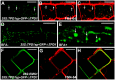
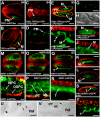
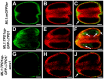
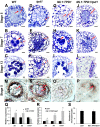
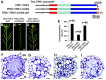
A fundamental feature of sexual reproduction in plants and animals is the specification of reproductive cells that conduct meiosis to form gametes, and the associated somatic cells that provide nutrition and developmental cues to ensure successful gamete production. The anther, which is the male reproductive organ in seed plants, produces reproductive microsporocytes (pollen mother cells) and surrounding somatic cells. The microsporocytes yield pollen via meiosis, and the somatic cells, particularly the tapetum, are required for the normal development of pollen. It is not known how the reproductive cells affect the differentiation of these somatic cells, and vice versa. Here, we use molecular genetics, cell biological, and biochemical approaches to demonstrate that TPD1 (TAPETUM DETERMINANT1) is a small secreted cysteine-rich protein ligand that interacts with the LRR (Leucine-Rich Repeat) domain of the EMS1 (EXCESS MICROSPOROCYTES1) receptor kinase at two sites. Analyses of the expressions and localizations of TPD1 and EMS1, ectopic expression of TPD1, experimental missorting of TPD1, and ablation of microsporocytes yielded results suggesting that the precursors of microsporocyte/microsporocyte-derived TPD1 and pre-tapetal-cell-localized EMS1 initially promote the periclinal division of secondary parietal cells and then determine one of the two daughter cells as a functional tapetal cell. Our results also indicate that tapetal cells suppress microsporocyte proliferation. Collectively, our findings show that tapetal cell differentiation requires reproductive-cell-secreted TPD1, illuminating a novel mechanism whereby signals from reproductive cells determine somatic cell fate in plant sexual reproduction.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Tapetum determinant1 is required for cell specialization in the Arabidopsis anther.Plant Cell. 2003 Dec;15(12):2792-804. doi: 10.1105/tpc.016618. Epub 2003 Nov 13. Plant Cell. 2003. PMID: 14615601 Free PMC article.
-
Ectopic expression of TAPETUM DETERMINANT1 affects ovule development in Arabidopsis.J Exp Bot. 2016 Mar;67(5):1311-26. doi: 10.1093/jxb/erv523. Epub 2015 Dec 17. J Exp Bot. 2016. PMID: 26685185
-
Overexpression of TAPETUM DETERMINANT1 alters the cell fates in the Arabidopsis carpel and tapetum via genetic interaction with excess microsporocytes1/extra sporogenous cells.Plant Physiol. 2005 Sep;139(1):186-91. doi: 10.1104/pp.105.063529. Epub 2005 Jul 29. Plant Physiol. 2005. PMID: 16055681 Free PMC article.
-
Control of anther cell differentiation: a teamwork of receptor-like kinases.Sex Plant Reprod. 2009 Dec;22(4):221-8. doi: 10.1007/s00497-009-0106-3. Epub 2009 Aug 6. Sex Plant Reprod. 2009. PMID: 20033443 Review.
-
Specification of tapetum and microsporocyte cells within the anther.Curr Opin Plant Biol. 2014 Feb;17:49-55. doi: 10.1016/j.pbi.2013.11.001. Epub 2013 Dec 3. Curr Opin Plant Biol. 2014. PMID: 24507494 Review.
Cited by
-
Sterility Caused by Floral Organ Degeneration and Abiotic Stresses in Arabidopsis and Cereal Grains.Front Plant Sci. 2016 Oct 14;7:1503. doi: 10.3389/fpls.2016.01503. eCollection 2016. Front Plant Sci. 2016. PMID: 27790226 Free PMC article. Review.
-
Tapetum-Dependent Male Meiosis Progression in Plants: Increasing Evidence Emerges.Front Plant Sci. 2020 Jan 16;10:1667. doi: 10.3389/fpls.2019.01667. eCollection 2019. Front Plant Sci. 2020. PMID: 32010157 Free PMC article. Review.
-
Rhizobium sp. IRBG74 Alters Arabidopsis Root Development by Affecting Auxin Signaling.Front Microbiol. 2018 Jan 4;8:2556. doi: 10.3389/fmicb.2017.02556. eCollection 2017. Front Microbiol. 2018. PMID: 29354099 Free PMC article.
-
Morphological Characterization of a New and Easily Recognizable Nuclear Male Sterile Mutant of Sorghum (Sorghum bicolor).PLoS One. 2017 Jan 4;12(1):e0165195. doi: 10.1371/journal.pone.0165195. eCollection 2017. PLoS One. 2017. PMID: 28052078 Free PMC article.
-
CIK Receptor Kinases Determine Cell Fate Specification during Early Anther Development in Arabidopsis.Plant Cell. 2018 Oct;30(10):2383-2401. doi: 10.1105/tpc.17.00586. Epub 2018 Sep 10. Plant Cell. 2018. PMID: 30201822 Free PMC article.
References
-
- Walbot V, Evans MM (2003) Unique features of the plant life cycle and their consequences. Nat Rev Genet 4: 369–379. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

