Tyrosinase-Cre-Mediated Deletion of the Autophagy Gene Atg7 Leads to Accumulation of the RPE65 Variant M450 in the Retinal Pigment Epithelium of C57BL/6 Mice
- PMID: 27537685
- PMCID: PMC4990303
- DOI: 10.1371/journal.pone.0161640
Tyrosinase-Cre-Mediated Deletion of the Autophagy Gene Atg7 Leads to Accumulation of the RPE65 Variant M450 in the Retinal Pigment Epithelium of C57BL/6 Mice
Abstract
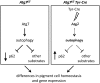
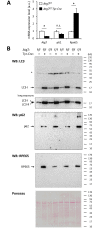
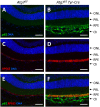
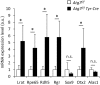
Targeted gene knockout mouse models have helped to identify roles of autophagy in many tissues. Here, we investigated the retinal pigment epithelium (RPE) of Atg7f/f Tyr-Cre mice (on a C57BL/6 background), in which Cre recombinase is expressed under the control of the tyrosinase promoter to delete the autophagy gene Atg7. In line with pigment cell-directed blockade of autophagy, the RPE and the melanocytes of the choroid showed strong accumulation of the autophagy adaptor and substrate, sequestosome 1 (Sqstm1)/p62, relative to the levels in control mice. Immunofluorescence and Western blot analysis demonstrated that the RPE, but not the choroid melanocytes, of Atg7f/f Tyr-Cre mice also had strongly increased levels of retinoid isomerohydrolase RPE65, a pivotal enzyme for the maintenance of visual perception. In contrast to Sqstm1, genes involved in retinal regeneration, i.e. Lrat, Rdh5, Rgr, and Rpe65, were expressed at higher mRNA levels. Sequencing of the Rpe65 gene showed that Atg7f/f and Atg7f/f Tyr-Cre mice carry a point mutation (L450M) that is characteristic for the C57BL/6 mouse strain and reportedly causes enhanced degradation of the RPE65 protein by an as-yet unknown mechanism. These results suggest that the increased abundance of RPE65 M450 in the RPE of Atg7f/f Tyr-Cre mice is, at least partly, mediated by upregulation of Rpe65 transcription; however, our data are also compatible with the hypothesis that the RPE65 M450 protein is degraded by Atg7-dependent autophagy in Atg7f/f mice. Further studies in mice of different genetic backgrounds are necessary to determine the relative contributions of these mechanisms.
Conflict of interest statement
Figures
Similar articles
-
Early AMD-like defects in the RPE and retinal degeneration in aged mice with RPE-specific deletion of Atg5 or Atg7.Mol Vis. 2017 Apr 14;23:228-241. eCollection 2017. Mol Vis. 2017. PMID: 28465655 Free PMC article.
-
Filamentous Aggregation of Sequestosome-1/p62 in Brain Neurons and Neuroepithelial Cells upon Tyr-Cre-Mediated Deletion of the Autophagy Gene Atg7.Mol Neurobiol. 2018 Nov;55(11):8425-8437. doi: 10.1007/s12035-018-0996-x. Epub 2018 Mar 17. Mol Neurobiol. 2018. PMID: 29550918 Free PMC article.
-
Somatic ablation of the Lrat gene in the mouse retinal pigment epithelium drastically reduces its retinoid storage.Invest Ophthalmol Vis Sci. 2007 Dec;48(12):5377-87. doi: 10.1167/iovs.07-0673. Invest Ophthalmol Vis Sci. 2007. PMID: 18055784
-
Retinal pigment epithelium 65 kDa protein (RPE65): An update.Prog Retin Eye Res. 2022 May;88:101013. doi: 10.1016/j.preteyeres.2021.101013. Epub 2021 Oct 2. Prog Retin Eye Res. 2022. PMID: 34607013 Free PMC article. Review.
-
RPE65 Palmitoylation: A Tale of Lipid Posttranslational Modification.Adv Exp Med Biol. 2019;1185:537-541. doi: 10.1007/978-3-030-27378-1_88. Adv Exp Med Biol. 2019. PMID: 31884667 Free PMC article. Review.
Cited by
-
Early AMD-like defects in the RPE and retinal degeneration in aged mice with RPE-specific deletion of Atg5 or Atg7.Mol Vis. 2017 Apr 14;23:228-241. eCollection 2017. Mol Vis. 2017. PMID: 28465655 Free PMC article.
-
HSP90 stabilizes visual cycle retinol dehydrogenase 5 in the endoplasmic reticulum by inhibiting its degradation during autophagy.J Biol Chem. 2025 Feb;301(2):108126. doi: 10.1016/j.jbc.2024.108126. Epub 2024 Dec 25. J Biol Chem. 2025. PMID: 39725039 Free PMC article.
-
Loss of CLN3, the gene mutated in juvenile neuronal ceroid lipofuscinosis, leads to metabolic impairment and autophagy induction in retinal pigment epithelium.Biochim Biophys Acta Mol Basis Dis. 2020 Oct 1;1866(10):165883. doi: 10.1016/j.bbadis.2020.165883. Epub 2020 Jun 25. Biochim Biophys Acta Mol Basis Dis. 2020. PMID: 32592935 Free PMC article.
-
The role and function of matrix metalloproteinase-8 in rhegmatogenous retinal detachment.Int J Clin Exp Pathol. 2017 Jul 1;10(7):7325-7332. eCollection 2017. Int J Clin Exp Pathol. 2017. PMID: 31966572 Free PMC article.
-
Autophagic Control of Skin Aging.Front Cell Dev Biol. 2019 Jul 30;7:143. doi: 10.3389/fcell.2019.00143. eCollection 2019. Front Cell Dev Biol. 2019. PMID: 31417903 Free PMC article. Review.
References
-
- Sukseree S, Eckhart L, Tschachler E, Watanapokasin R. Autophagy in epithelial homeostasis and defense. Front Biosci. 2013;E5:1000–1010. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

