Mutations in genes encoding antibiotic substances increase the synthesis of poly-γ-glutamic acid in Bacillus amyloliquefaciens LL3
- PMID: 27539744
- PMCID: PMC5300885
- DOI: 10.1002/mbo3.398
Mutations in genes encoding antibiotic substances increase the synthesis of poly-γ-glutamic acid in Bacillus amyloliquefaciens LL3
Abstract
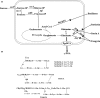
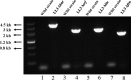
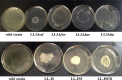
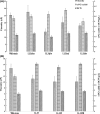
Poly-γ-glutamic acid (γ-PGA) is an important natural biopolymer that is used widely in fields of foods, medicine, cosmetics, and agriculture. Several B. amyloliquefaciens LL3 mutants were constructed to improve γ-PGA synthesis via single or multiple marker-less in-frame deletions of four gene clusters (itu, bae, srf, and fen) encoding antibiotic substances. γ-PGA synthesis by the Δsrf mutant showed a slight increase (4.1 g/L) compared with that of the wild-type strain (3.3 g/L). The ΔituΔsrf mutant showed increased γ-PGA yield from 3.3 to 4.5 g/L, with an increase of 36.4%. The γ-PGA yield of the ΔituΔsrfΔfen and ΔituΔsrfΔfenΔbae mutants did not show a further increase. The four gene clusters' roles in swarming motility and biofilm formation were also studied. The Δsrf and Δbae mutant strains were both significantly defective in swarming, indicating that bacillaene and surfactin are involved in swarming motility of B. amyloliquefaciens LL3. Furthermore, Δsrf and Δitu mutant strains were obviously defective in biofilm formation; therefore, iturin and surfactin must play important roles in biofilm formation in B. amyloliquefaciens LL3.
Keywords: antibiotic substance; biofilm formation; gene marker-less deletion; poly-γ-glutamic acid; swarming.
© 2016 The Authors. MicrobiologyOpen published by John Wiley & Sons Ltd.
Figures

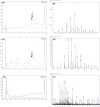
Similar articles
-
Metabolic engineering of Bacillus amyloliquefaciens LL3 for enhanced poly-γ-glutamic acid synthesis.Microb Biotechnol. 2019 Sep;12(5):932-945. doi: 10.1111/1751-7915.13446. Epub 2019 Jun 20. Microb Biotechnol. 2019. PMID: 31219230 Free PMC article.
-
Effects of MreB paralogs on poly-γ-glutamic acid synthesis and cell morphology in Bacillus amyloliquefaciens.FEMS Microbiol Lett. 2016 Sep;363(17):fnw187. doi: 10.1093/femsle/fnw187. Epub 2016 Jul 31. FEMS Microbiol Lett. 2016. PMID: 27481703
-
Deletion of genes involved in glutamate metabolism to improve poly-gamma-glutamic acid production in B. amyloliquefaciens LL3.J Ind Microbiol Biotechnol. 2015 Feb;42(2):297-305. doi: 10.1007/s10295-014-1563-8. Epub 2014 Dec 25. J Ind Microbiol Biotechnol. 2015. PMID: 25540046
-
Genetic and metabolic engineering for microbial production of poly-γ-glutamic acid.Biotechnol Adv. 2018 Sep-Oct;36(5):1424-1433. doi: 10.1016/j.biotechadv.2018.05.006. Epub 2018 May 28. Biotechnol Adv. 2018. PMID: 29852203 Review.
-
New biological functions and applications of high-molecular-mass poly-gamma-glutamic acid.Chem Biodivers. 2010 Jun;7(6):1555-62. doi: 10.1002/cbdv.200900283. Chem Biodivers. 2010. PMID: 20564573 Review.
Cited by
-
Baicalein Inhibits the Progression and Promotes Radiosensitivity of Esophageal Squamous Cell Carcinoma by Targeting HIF-1A.Drug Des Devel Ther. 2022 Jul 29;16:2423-2436. doi: 10.2147/DDDT.S370114. eCollection 2022. Drug Des Devel Ther. 2022. PMID: 35937565 Free PMC article.
-
Metabolic engineering of Bacillus amyloliquefaciens LL3 for enhanced poly-γ-glutamic acid synthesis.Microb Biotechnol. 2019 Sep;12(5):932-945. doi: 10.1111/1751-7915.13446. Epub 2019 Jun 20. Microb Biotechnol. 2019. PMID: 31219230 Free PMC article.
-
Antagonism of Two Plant-Growth Promoting Bacillus velezensis Isolates Against Ralstonia solanacearum and Fusarium oxysporum.Sci Rep. 2018 Mar 12;8(1):4360. doi: 10.1038/s41598-018-22782-z. Sci Rep. 2018. PMID: 29531357 Free PMC article.
-
Engineering of a genome-reduced strain Bacillus amyloliquefaciens for enhancing surfactin production.Microb Cell Fact. 2020 Dec 7;19(1):223. doi: 10.1186/s12934-020-01485-z. Microb Cell Fact. 2020. PMID: 33287813 Free PMC article.
-
Enhanced production of antifungal lipopeptide iturin A by Bacillus amyloliquefaciens LL3 through metabolic engineering and culture conditions optimization.Microb Cell Fact. 2019 Apr 10;18(1):68. doi: 10.1186/s12934-019-1121-1. Microb Cell Fact. 2019. PMID: 30971238 Free PMC article.
References
-
- Abdel‐Mawgoud, A. M. , Aboulwafa, M. M. , & Hassouna, N. A. H. (2008). Optimization of surfactin production by Bacillus subtilis isolate BS5. Applied Biochemistry and Biotechnology, 150, 305–325. - PubMed
-
- Cao, M. F. , Geng, W. T. , Liu, L. , Song, C. J. , Xie, H. , Guo, W. B. , … Wang, S. F. (2011). Glutamic acid independent production of poly‐γ‐glutamic acid by LL3 and cloning of pgsBCA genes. Bioresource Technology, 102, 4251–4257. - PubMed
-
- Chen, X. H. , Koumoutsi, A. , & Scholz, R. (2009). More than anticipated ‐ production of antibiotics and other secondary metabolites by Bacillus amyloliquefaciens FZB42. Journal of Molecular Microbiology and Biotechnology, 16, 14–24. - PubMed
-
- Chen, X. H. , Koumoutsia, A. , Scholza, R. , Schneiderb, K. , Vaterb, J. , Süssmuthb, R. , … Borrissa, R. (2009). Genome analysis of Bacillus amyloliquefaciens FZB42 reveals its potential for biocontrol of plant pathogens. Journal of Biotechnology, 140, 27–37. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

