Metabolic requirements for cancer cell proliferation
- PMID: 27540483
- PMCID: PMC4989334
- DOI: 10.1186/s40170-016-0156-6
Metabolic requirements for cancer cell proliferation
Abstract
Background: The study of cancer metabolism has been largely dedicated to exploring the hypothesis that oncogenic transformation rewires cellular metabolism to sustain elevated rates of growth and division. Intense examination of tumors and cancer cell lines has confirmed that many cancer-associated metabolic phenotypes allow robust growth and survival; however, little attention has been given to explicitly identifying the biochemical requirements for cell proliferation in a rigorous manner in the context of cancer metabolism.
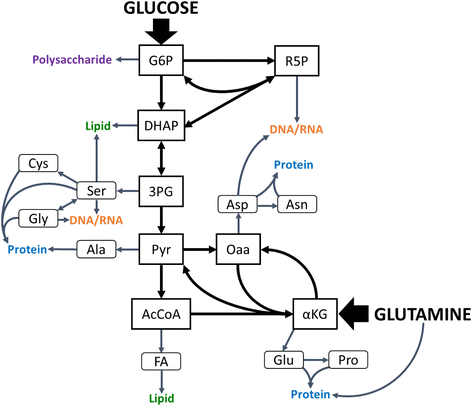
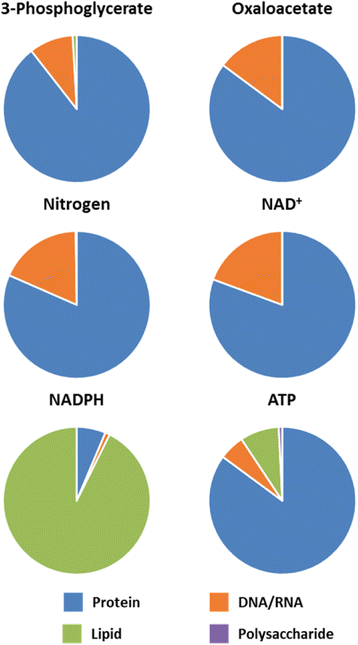
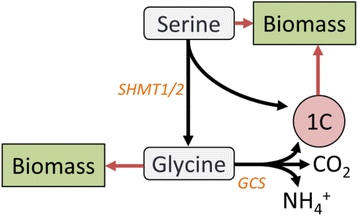
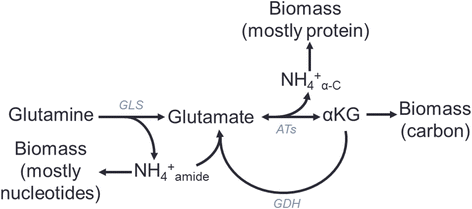
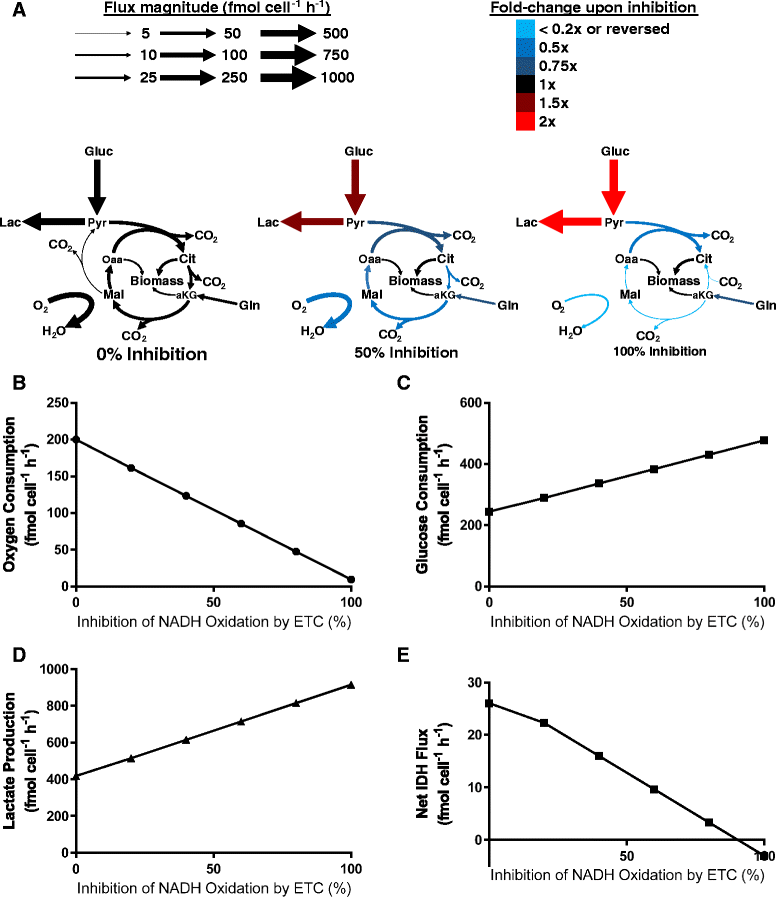
Results: Using a well-studied hybridoma line as a model, we comprehensively and quantitatively enumerate the metabolic requirements for generating new biomass in mammalian cells; this indicated a large biosynthetic requirement for ATP, NADPH, NAD(+), acetyl-CoA, and amino acids. Extension of this approach to serine/glycine and glutamine metabolic pathways suggested lower limits on serine and glycine catabolism to supply one-carbon unit synthesis and significant availability of glutamine-derived carbon for biosynthesis resulting from nitrogen demands alone, respectively. We integrated our biomass composition results into a flux balance analysis model, placing upper bounds on mitochondrial NADH oxidation to simulate metformin treatment; these simulations reproduced several empirically observed metabolic phenotypes, including increased reductive isocitrate dehydrogenase flux.
Conclusions: Our analysis clarifies the differential needs for central carbon metabolism precursors, glutamine-derived nitrogen, and cofactors such as ATP, NADPH, and NAD(+), while also providing justification for various extracellular nutrient uptake behaviors observed in tumors. Collectively, these results demonstrate how stoichiometric considerations alone can successfully predict empirically observed phenotypes and provide insight into biochemical dynamics that underlie responses to metabolic perturbations.
Keywords: Anabolism; Biosynthesis; Cancer metabolism; Metabolic modeling; Proliferation; Stoichiometric analysis.
Figures
References
-
- Hitosugi T, Kang S, Vander Heiden MG, Chung T-W, Elf S, Lythgoe K, Dong S, Lonial S, Wang X, Chen GZ, Xie J, Gu T-L, Polakiewicz RD, Roesel JL, Boggon TJ, Khuri FR, Gilliland DG, Cantley LC, Kaufman J, Chen J. Tyrosine phosphorylation inhibits PKM2 to promote the Warburg effect and tumor growth. Sci Signal. 2009;2:ra73. doi: 10.1126/scisignal.2000431. - DOI - PMC - PubMed
-
- Vander Heiden MG, Lunt SY, Dayton TL, Fiske BP, Israelsen WJ, Mattaini KR, Vokes NI, Stephanopoulos G, Cantley LC, Metallo CM, Locasale JW: Metabolic pathway alterations that support cell proliferation. Cold Spring Harb Symp Quant Biol. 2012;76:325-34. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

