A reference panel of 64,976 haplotypes for genotype imputation
- PMID: 27548312
- PMCID: PMC5388176
- DOI: 10.1038/ng.3643
A reference panel of 64,976 haplotypes for genotype imputation
Abstract
We describe a reference panel of 64,976 human haplotypes at 39,235,157 SNPs constructed using whole-genome sequence data from 20 studies of predominantly European ancestry. Using this resource leads to accurate genotype imputation at minor allele frequencies as low as 0.1% and a large increase in the number of SNPs tested in association studies, and it can help to discover and refine causal loci. We describe remote server resources that allow researchers to carry out imputation and phasing consistently and efficiently.
Figures
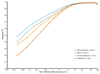

References
-
- Genome of the Netherlands Consortium. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat Genet. 2014;46:818–825. - PubMed
Publication types
MeSH terms
Grants and funding
- R01 CA059045/CA/NCI NIH HHS/United States
- U01 DK085501/DK/NIDDK NIH HHS/United States
- R01 DK098032/DK/NIDDK NIH HHS/United States
- WT091310/WT_/Wellcome Trust/United Kingdom
- R01 MH094145/MH/NIMH NIH HHS/United States
- UM1 CA167552/CA/NCI NIH HHS/United States
- R01 MH085548/MH/NIMH NIH HHS/United States
- R01 DK093757/DK/NIDDK NIH HHS/United States
- HHSN268201100001C/WH/WHI NIH HHS/United States
- G0601966/MRC_/Medical Research Council/United Kingdom
- T32 GM007753/GM/NIGMS NIH HHS/United States
- HHSN268201300049C/HL/NHLBI NIH HHS/United States
- U54 DK083912/DK/NIDDK NIH HHS/United States
- CZB/4/710/CSO_/Chief Scientist Office/United Kingdom
- U01 DK085545/DK/NIDDK NIH HHS/United States
- HHSN268201300046C/HL/NHLBI NIH HHS/United States
- R01 CA137178/CA/NCI NIH HHS/United States
- ZIA EY000546/EY/NEI NIH HHS/United States
- G0900747/MRC_/Medical Research Council/United Kingdom
- R01 MH101820/MH/NIMH NIH HHS/United States
- WT097307/WT_/Wellcome Trust/United Kingdom
- MC_UU_12013/3/MRC_/Medical Research Council/United Kingdom
- SP/04/002/BHF_/British Heart Foundation/United Kingdom
- U01 CA137088/CA/NCI NIH HHS/United States
- U19 AG023122/AG/NIA NIH HHS/United States
- R01 MH090937/MH/NIMH NIH HHS/United States
- R01 DK072193/DK/NIDDK NIH HHS/United States
- K24 DK080140/DK/NIDDK NIH HHS/United States
- R01 AA009367/AA/NIAAA NIH HHS/United States
- HHSN271201100004C/AG/NIA NIH HHS/United States
- HHSN268201300047C/HL/NHLBI NIH HHS/United States
- R01 CA151993/CA/NCI NIH HHS/United States
- R01 HG000376/HG/NHGRI NIH HHS/United States
- U01 MH105653/MH/NIMH NIH HHS/United States
- N01 MH080001/MH/NIMH NIH HHS/United States
- R01 DA036216/DA/NIDA NIH HHS/United States
- RC2 DK088389/DK/NIDDK NIH HHS/United States
- R01 DK073541/DK/NIDDK NIH HHS/United States
- HHSN268201100004C/WH/WHI NIH HHS/United States
- MC_PC_15018/MRC_/Medical Research Council/United Kingdom
- R01 MH104964/MH/NIMH NIH HHS/United States
- HHSN268201300048C/HL/NHLBI NIH HHS/United States
- WT090851/WT_/Wellcome Trust/United Kingdom
- ARC_/Arthritis Research UK/United Kingdom
- U01 DK085526/DK/NIDDK NIH HHS/United States
- MR/J00314X/1/MRC_/Medical Research Council/United Kingdom
- U01 HG005773/HG/NHGRI NIH HHS/United States
- U01 DK062370/DK/NIDDK NIH HHS/United States
- HHSN268201100046C/HL/NHLBI NIH HHS/United States
- P30 EY001583/EY/NEI NIH HHS/United States
- HHSN268201100003C/WH/WHI NIH HHS/United States
- R01 DA037904/DA/NIDA NIH HHS/United States
- CZB/4/276/CSO_/Chief Scientist Office/United Kingdom
- N01EY50007/EY/NEI NIH HHS/United States
- R01 HL127564/HL/NHLBI NIH HHS/United States
- R01 HG007022/HG/NHGRI NIH HHS/United States
- K99 DK092251/DK/NIDDK NIH HHS/United States
- U01 CA164930/CA/NCI NIH HHS/United States
- U01 HL117626/HL/NHLBI NIH HHS/United States
- HHSN268201100002C/WH/WHI NIH HHS/United States
- U01 MH105641/MH/NIMH NIH HHS/United States
- R01 HL117626/HL/NHLBI NIH HHS/United States
- P01 CA055075/CA/NCI NIH HHS/United States
- R01 DA024417/DA/NIDA NIH HHS/United States
- HHSN268201300050C/HL/NHLBI NIH HHS/United States
- R01 HL109946/HL/NHLBI NIH HHS/United States
- MOP-172013/CIHR/Canada
- 202802/Z/16/Z/WT_/Wellcome Trust/United Kingdom
- P50 CA127003/CA/NCI NIH HHS/United States
- G0601261/MRC_/Medical Research Council/United Kingdom
- R01 HL102830/HL/NHLBI NIH HHS/United States
- R01 DA005147/DA/NIDA NIH HHS/United States
- WT098051/WT_/Wellcome Trust/United Kingdom
- P30 DK020595/DK/NIDDK NIH HHS/United States
- G9815508/MRC_/Medical Research Council/United Kingdom
- G0700931/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources

