Spatiotemporal Fluctuation Analysis: A Powerful Tool for the Future Nanoscopy of Molecular Processes
- PMID: 27558712
- PMCID: PMC5002078
- DOI: 10.1016/j.bpj.2016.07.015
Spatiotemporal Fluctuation Analysis: A Powerful Tool for the Future Nanoscopy of Molecular Processes
Abstract
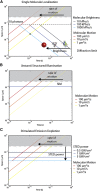
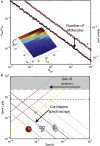
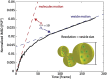
The enormous wealth of information available today from optical microscopy measurements on living samples is often underexploited. We argue that spatiotemporal analysis of fluorescence fluctuations using multiple detection channels can enhance the performance of current nanoscopy methods and provide further insight into dynamic molecular processes of high biological relevance.
Copyright © 2016 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
Similar articles
-
Zooming in on biological processes with fluorescence nanoscopy.Curr Opin Biotechnol. 2013 Aug;24(4):646-53. doi: 10.1016/j.copbio.2013.02.016. Epub 2013 Mar 13. Curr Opin Biotechnol. 2013. PMID: 23498844 Review.
-
A new wave of cellular imaging.Annu Rev Cell Dev Biol. 2010;26:285-314. doi: 10.1146/annurev-cellbio-100109-104048. Annu Rev Cell Dev Biol. 2010. PMID: 20929313 Review.
-
Fast stimulated emission nanoscopy based on single molecule localization.Appl Opt. 2015 Aug 1;54(22):6919-23. doi: 10.1364/AO.54.006919. Appl Opt. 2015. PMID: 26368110
-
Multicolor fluorescence nanoscopy by photobleaching: concept, verification, and its application to resolve selective storage of proteins in platelets.ACS Nano. 2014 May 27;8(5):4358-65. doi: 10.1021/nn406113m. Epub 2014 Apr 24. ACS Nano. 2014. PMID: 24730587
-
Live-Cell Super-resolution Fluorescence Microscopy.Biochemistry (Mosc). 2019 Jan;84(Suppl 1):S19-S31. doi: 10.1134/S0006297919140025. Biochemistry (Mosc). 2019. PMID: 31213193 Review.
Cited by
-
Spatiotemporal Correlation Spectroscopy Reveals a Protective Effect of Peptide-Based GLP-1 Receptor Agonism against Lipotoxicity on Insulin Granule Dynamics in Primary Human β-Cells.Pharmaceutics. 2021 Sep 3;13(9):1403. doi: 10.3390/pharmaceutics13091403. Pharmaceutics. 2021. PMID: 34575477 Free PMC article.
-
Quantitative Bio-Imaging Tools to Dissect the Interplay of Membrane and Cytoskeletal Actin Dynamics in Immune Cells.Front Immunol. 2021 Jan 11;11:612542. doi: 10.3389/fimmu.2020.612542. eCollection 2020. Front Immunol. 2021. PMID: 33505401 Free PMC article. Review.
-
Time-resolved biophysical approaches to nucleocytoplasmic transport.Comput Struct Biotechnol J. 2017 Apr 4;15:299-306. doi: 10.1016/j.csbj.2017.03.005. eCollection 2017. Comput Struct Biotechnol J. 2017. PMID: 28435614 Free PMC article. Review.
-
Anomalous Diffusion in Inverted Variable-Lengthscale Fluorescence Correlation Spectroscopy.Biophys J. 2019 Mar 5;116(5):791-806. doi: 10.1016/j.bpj.2019.01.024. Epub 2019 Jan 30. Biophys J. 2019. PMID: 30782396 Free PMC article.
-
Pair correlation microscopy of intracellular molecular transport.Nat Protoc. 2025 Jun;20(6):1651-1677. doi: 10.1038/s41596-024-01097-6. Epub 2025 Feb 6. Nat Protoc. 2025. PMID: 39915621 Review.
References
-
- Hell S.W., Sahl S.J., Tinnefeld P. The 2015 super-resolution microscopy roadmap. J. Phys. D Appl. Phys. 2015;48:443001.
-
- Abbe E. Beiträge zur Theorie des Mikroskops und der mikroskopischen Wahrnehmung. Arch. F. Microsc. Anat. 1873;9:413–418.
-
- Hell S.W., Wichmann J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 1994;19:780–782. - PubMed
-
- Hell S.W., Kroug M. Ground-state-depletion fluorscence microscopy: a concept for breaking the diffraction resolution limit. Appl. Phys. B. 1995;60:495–497.
-
- Hell S., Jakobs S., Kastrup L. Imaging and writing at the nanoscale with focused visible light through saturable optical transitions. Appl. Phys. A Mater. Sci. Process. 2003;77:859–860.
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

