Pseudogenes regulate parental gene expression via ceRNA network
- PMID: 27561207
- PMCID: PMC5192809
- DOI: 10.1111/jcmm.12952
Pseudogenes regulate parental gene expression via ceRNA network
Abstract
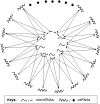
The concept of competitive endogenous RNA (ceRNA) was first proposed by Salmena and colleagues. Evidence suggests that pseudogene RNAs can act as a 'sponge' through competitive binding of common miRNA, releasing or attenuating repression through sequestering miRNAs away from parental mRNA. In theory, ceRNAs refer to all transcripts such as mRNA, tRNA, rRNA, long non-coding RNA, pseudogene RNA and circular RNA, because all of them may become the targets of miRNA depending on spatiotemporal situation. As binding of miRNA to the target RNA is not 100% complementary, it is possible that one miRNA can bind to multiple target RNAs and vice versa. All RNAs crosstalk through competitively binding to miRNAvia miRNA response elements (MREs) contained within the RNA sequences, thus forming a complex regulatory network. The ratio of a subset of miRNAs to the corresponding number of MREs determines repression strength on a given mRNA translation or stability. An increase in pseudogene RNA level can sequester miRNA and release repression on the parental gene, leading to an increase in parental gene expression. A massive number of transcripts constitute a complicated network that regulates each other through this proposed mechanism, though some regulatory significance may be mild or even undetectable. It is possible that the regulation of gene and pseudogene expression occurring in this manor involves all RNAs bearing common MREs. In this review, we will primarily discuss how pseudogene transcripts regulate expression of parental genes via ceRNA network and biological significance of regulation.
Keywords: ceRNA; gene expression; microRNA; pseudogene.
© 2016 The Authors. Journal of Cellular and Molecular Medicine published by John Wiley & Sons Ltd and Foundation for Cellular and Molecular Medicine.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

