TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription
- PMID: 27564129
- PMCID: PMC5053588
- DOI: 10.1080/15384101.2016.1222337
TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription
Abstract
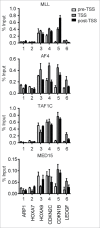
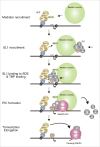
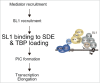
Gene rearrangement of the mixed lineage leukemia (MLL) gene causes leukemia by inducing the constitutive expression of a gene subset normally expressed only in the immature haematopoietic progenitor cells. MLL gene rearrangements often generate fusion products of MLL and a component of the AF4 family/ENL family/P-TEFb (AEP) complex. MLL-AEP fusion proteins have the potential of constitutively recruiting the P-TEFb elongation complex. Thus, it is hypothesized that relieving the promoter proximal pausing of RNA polymerase II is the rate-limiting step of MLL fusion-dependent transcription. AEP also has the potential to recruit the mediator complex via MED26. We recently showed that AEP activates transcription initiation by facilitating TBP loading to the TATA element through the SL1 complex. In the present study, we show that the key activity responsible for the oncogenic property of MLL-AEP fusion proteins is the TBP loading activity, and not the mediator recruitment or transcriptional elongation activities. Thus, we propose that TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription.
Keywords: AF4; MLL; SL1; TBP; leukemia; mediator; transcription.
Figures



Similar articles
-
AF4 uses the SL1 components of RNAP1 machinery to initiate MLL fusion- and AEP-dependent transcription.Nat Commun. 2015 Nov 23;6:8869. doi: 10.1038/ncomms9869. Nat Commun. 2015. PMID: 26593443 Free PMC article.
-
Transcriptional activation by MLL fusion proteins in leukemogenesis.Exp Hematol. 2017 Feb;46:21-30. doi: 10.1016/j.exphem.2016.10.014. Epub 2016 Nov 16. Exp Hematol. 2017. PMID: 27865805 Review.
-
RNA Polymerase II-Dependent Transcription Initiated by Selectivity Factor 1: A Central Mechanism Used by MLL Fusion Proteins in Leukemic Transformation.Front Genet. 2019 Jan 14;9:722. doi: 10.3389/fgene.2018.00722. eCollection 2018. Front Genet. 2019. PMID: 30693017 Free PMC article. Review.
-
A higher-order complex containing AF4 and ENL family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription.Cancer Cell. 2010 Feb 17;17(2):198-212. doi: 10.1016/j.ccr.2009.12.040. Cancer Cell. 2010. PMID: 20153263 Free PMC article.
-
Cooperative gene activation by AF4 and DOT1L drives MLL-rearranged leukemia.J Clin Invest. 2017 May 1;127(5):1918-1931. doi: 10.1172/JCI91406. Epub 2017 Apr 10. J Clin Invest. 2017. PMID: 28394257 Free PMC article.
Cited by
-
HBO1-MLL interaction promotes AF4/ENL/P-TEFb-mediated leukemogenesis.Elife. 2021 Aug 25;10:e65872. doi: 10.7554/eLife.65872. Elife. 2021. PMID: 34431785 Free PMC article.
-
RNA-binding proteins of KHDRBS and IGF2BP families control the oncogenic activity of MLL-AF4.Nat Commun. 2022 Nov 5;13(1):6688. doi: 10.1038/s41467-022-34558-1. Nat Commun. 2022. PMID: 36335100 Free PMC article.
-
Modeling by disruption and a selected-for partner for the nude locus.EMBO Rep. 2021 Mar 3;22(3):e49804. doi: 10.15252/embr.201949804. Epub 2020 Dec 28. EMBO Rep. 2021. PMID: 33369874 Free PMC article.
-
Molecular processes involved in B cell acute lymphoblastic leukaemia.Cell Mol Life Sci. 2018 Feb;75(3):417-446. doi: 10.1007/s00018-017-2620-z. Epub 2017 Aug 17. Cell Mol Life Sci. 2018. PMID: 28819864 Free PMC article. Review.
-
Another piece of the puzzle added to understand t(4;11) leukemia better.Haematologica. 2019 Jun;104(6):1098-1100. doi: 10.3324/haematol.2018.213397. Haematologica. 2019. PMID: 31152087 Free PMC article. No abstract available.
References
-
- Basehoar AD, Zanton SJ, Pugh BF. Identification and distinct regulation of yeast TATA box-containing genes. Cell 2004; 116:699-709; PMID:15006352; http://dx.doi.org/10.1016/S0092-8674(04)00205-3 - DOI - PubMed
-
- Rhee HS, Pugh BF. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature 2012; 483:295-301; PMID:22258509; http://dx.doi.org/10.1038/nature10799 - DOI - PMC - PubMed
-
- Nilson I, Reichel M, Ennas MG, Greim R, Knorr C, Siegler G, Greil J, Fey GH, Marschalek R. Exon/intron structure of the human AF-4 gene, a member of the AF-4/LAF-4/FMR-2 gene family coding for a nuclear protein with structural alterations in acute leukaemia. Br J Haematol 1997; 98:157-69; PMID:9233580; http://dx.doi.org/10.1046/j.1365-2141.1997.1522966.x - DOI - PubMed
-
- Estable MC, Naghavi MH, Kato H, Xiao H, Qin J, Vahlne A, Roeder RG. MCEF, the newest member of the AF4 family of transcription factors involved in leukemia, is a positive transcription elongation factor-b-associated protein. J Biomed Sci 2002; 9:234-45; PMID:12065898; http://dx.doi.org/10.1007/BF02256070 - DOI - PubMed
-
- Benedikt A, Baltruschat S, Scholz B, Bursen A, Arrey TN, Meyer B, Varagnolo L, Muller AM, Karas M, Dingermann T, et al.. The leukemogenic AF4-MLL fusion protein causes P-TEFb kinase activation and altered epigenetic signatures. Leukemia 2011; 25:135-44; PMID:21030982; http://dx.doi.org/10.1038/leu.2010.249 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
