RB1 deficiency in triple-negative breast cancer induces mitochondrial protein translation
- PMID: 27571409
- PMCID: PMC5096803
- DOI: 10.1172/JCI81568
RB1 deficiency in triple-negative breast cancer induces mitochondrial protein translation
Abstract
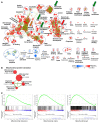
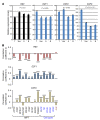
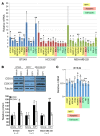
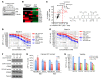
Triple-negative breast cancer (TNBC) includes basal-like and claudin-low subtypes for which no specific treatment is currently available. Although the retinoblastoma tumor-suppressor gene (RB1) is frequently lost together with TP53 in TNBC, it is not directly targetable. There is thus great interest in identifying vulnerabilities downstream of RB1 that can be therapeutically exploited. Here, we determined that combined inactivation of murine Rb and p53 in diverse mammary epithelial cells induced claudin-low-like TNBC with Met, Birc2/3-Mmp13-Yap1, and Pvt1-Myc amplifications. Gene set enrichment analysis revealed that Rb/p53-deficient tumors showed elevated expression of the mitochondrial protein translation (MPT) gene pathway relative to tumors harboring p53 deletion alone. Accordingly, bioinformatic, functional, and biochemical analyses showed that RB1-E2F complexes bind to MPT gene promoters to regulate transcription and control MPT. Additionally, a screen of US Food and Drug Administration-approved (FDA-approved) drugs identified the MPT antagonist tigecycline (TIG) as a potent inhibitor of Rb/p53-deficient tumor cell proliferation. TIG preferentially suppressed RB1-deficient TNBC cell proliferation, targeted both the bulk and cancer stem cell fraction, and strongly attenuated xenograft growth. It also cooperated with sulfasalazine, an FDA-approved inhibitor of cystine xCT antiporter, in culture and xenograft assays. Our results suggest that RB1 deficiency promotes cancer cell proliferation in part by enhancing mitochondrial function and identify TIG as a clinically approved drug for RB1-deficient TNBC.
Figures





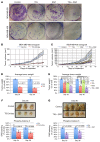

Similar articles
-
Targeted Pten deletion plus p53-R270H mutation in mouse mammary epithelium induces aggressive claudin-low and basal-like breast cancer.Breast Cancer Res. 2016 Jan 19;18(1):9. doi: 10.1186/s13058-015-0668-y. Breast Cancer Res. 2016. PMID: 26781438 Free PMC article.
-
RB1 status in triple negative breast cancer cells dictates response to radiation treatment and selective therapeutic drugs.PLoS One. 2013 Nov 12;8(11):e78641. doi: 10.1371/journal.pone.0078641. eCollection 2013. PLoS One. 2013. PMID: 24265703 Free PMC article.
-
RB loss contributes to aggressive tumor phenotypes in MYC-driven triple negative breast cancer.Cell Cycle. 2015;14(1):109-22. doi: 10.4161/15384101.2014.967118. Cell Cycle. 2015. PMID: 25602521 Free PMC article.
-
Mitochondrial OXPHOS Induced by RB1 Deficiency in Breast Cancer: Implications for Anabolic Metabolism, Stemness, and Metastasis.Trends Cancer. 2017 Nov;3(11):768-779. doi: 10.1016/j.trecan.2017.09.002. Epub 2017 Oct 17. Trends Cancer. 2017. PMID: 29120753 Review.
-
RB1 and p53 at the crossroad of EMT and triple-negative breast cancer.Cell Cycle. 2011 May 15;10(10):1563-70. doi: 10.4161/cc.10.10.15703. Epub 2011 May 15. Cell Cycle. 2011. PMID: 21502814 Review.
Cited by
-
CDC25 as a common therapeutic target for triple-negative breast cancer - the challenges ahead.Mol Cell Oncol. 2018 Aug 15;5(4):e1481814. doi: 10.1080/23723556.2018.1481814. eCollection 2018. Mol Cell Oncol. 2018. PMID: 30250928 Free PMC article.
-
Cross-species identification of PIP5K1-, splicing- and ubiquitin-related pathways as potential targets for RB1-deficient cells.PLoS Genet. 2021 Feb 16;17(2):e1009354. doi: 10.1371/journal.pgen.1009354. eCollection 2021 Feb. PLoS Genet. 2021. PMID: 33591981 Free PMC article.
-
Pan-cancer molecular analysis of the RB tumor suppressor pathway.Commun Biol. 2020 Apr 2;3(1):158. doi: 10.1038/s42003-020-0873-9. Commun Biol. 2020. PMID: 32242058 Free PMC article.
-
Identification of CDC25 as a Common Therapeutic Target for Triple-Negative Breast Cancer.Cell Rep. 2018 Apr 3;23(1):112-126. doi: 10.1016/j.celrep.2018.03.039. Cell Rep. 2018. PMID: 29617654 Free PMC article.
-
Intersection of retinoblastoma tumor suppressor function, stem cells, metabolism, and inflammation.Cancer Sci. 2017 Sep;108(9):1726-1731. doi: 10.1111/cas.13312. Epub 2017 Jul 29. Cancer Sci. 2017. PMID: 28865172 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

