Multi-omics analysis of primary glioblastoma cell lines shows recapitulation of pivotal molecular features of parental tumors
- PMID: 27571888
- PMCID: PMC5463853
- DOI: 10.1093/neuonc/now160
Multi-omics analysis of primary glioblastoma cell lines shows recapitulation of pivotal molecular features of parental tumors
Abstract
Background: Glioblastoma (GBM) is the deadliest primary brain cancer in adults. Emerging innovative therapies hold promise for personalized cancer treatment. Improving therapeutic options depends on research relying on relevant preclinical models. In this line we have established in the setting of the GlioTex project (GBM and Experimental Therapeutics) a GBM patient-derived cell line (GBM-PDCL) library. A multi-omic approach was used to determine the molecular landscape of PDCL and the extent to which they represent GBM tumors.
Methods: Single nucleotide polymorphism array, expression arrays, exome sequencing, and RNA sequencing were used to measure and compare the molecular landscapes of 20 samples representing 10 human GBM tumors and paired GBM-PDCLs.
Results: Copy number variations were similar for a median of 85% of the genome and for 59% of the major focal events. Somatic point mutations were similar in a median of 41%. Mutations in GBM driver and "druggable" genes were maintained in 67% of events. Mutations that were not conserved in the PDCL were mainly low allelic fraction and/or non-driver mutations. Based on RNA expression profiling, PDCLs cluster closely to their parental tumor with overexpression of pathways associated with cancer progression in PDCL.
Conclusions: Overall, PDCLs recapitulate pivotal molecular alterations of paired-parental tumors supporting their use as a preclinical model of GBM. However, some driver aberrations are lost or gained in the passage from tumor to PDCL. Our results support using PDCL as a relevant preclinical model of GBM. Further investigations of changes between PDCLs and their parental tumor may provide insights into GBM biology.
Keywords: cancer; cell lines; genome; glioblastoma.
© The Author(s) 2016. Published by Oxford University Press on behalf of the Society for Neuro-Oncology. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures

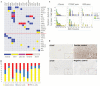

References
-
- Ciardiello F, Tortora G. EGFR antagonists in cancer treatment. N Engl J Med. 2008;358(11):1160–1174. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

