Virus-derived DNA drives mosquito vector tolerance to arboviral infection
- PMID: 27580708
- PMCID: PMC5025746
- DOI: 10.1038/ncomms12410
Virus-derived DNA drives mosquito vector tolerance to arboviral infection
Abstract
Mosquitoes develop long-lasting viral infections without substantial deleterious effects, despite high viral loads. This makes mosquitoes efficient vectors for emerging viral diseases with enormous burden on public health. How mosquitoes resist and/or tolerate these viruses is poorly understood. Here we show that two species of Aedes mosquitoes infected with two arboviruses from distinct families (dengue or chikungunya) generate a viral-derived DNA (vDNA) that is essential for mosquito survival and viral tolerance. Inhibition of vDNA formation leads to extreme susceptibility to viral infections, reduction of viral small RNAs due to an impaired immune response, and loss of viral tolerance. Our results highlight an essential role of vDNA in viral tolerance that allows mosquito survival and thus may be important for arbovirus dissemination and transmission. Elucidating the mechanisms of mosquito tolerance to arbovirus infection paves the way to conceptualize new antivectorial strategies to selectively eliminate arbovirus-infected mosquitoes.
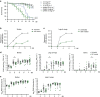
Figures



Similar articles
-
Entomological characterization of Aedes mosquitoes and arbovirus detection in Ibagué, a Colombian city with co-circulation of Zika, dengue and chikungunya viruses.Parasit Vectors. 2021 Sep 6;14(1):446. doi: 10.1186/s13071-021-04908-x. Parasit Vectors. 2021. PMID: 34488857 Free PMC article.
-
Profile of Small RNAs, vDNA Forms and Viral Integrations in Late Chikungunya Virus Infection of Aedes albopictus Mosquitoes.Viruses. 2021 Mar 25;13(4):553. doi: 10.3390/v13040553. Viruses. 2021. PMID: 33806250 Free PMC article.
-
No evidence for sylvatic cycles of chikungunya, dengue and Zika viruses in African green monkeys (Chlorocebus aethiops sabaeus) on St. Kitts, West Indies.Parasit Vectors. 2020 Oct 30;13(1):540. doi: 10.1186/s13071-020-04419-1. Parasit Vectors. 2020. PMID: 33126907 Free PMC article.
-
Human Urban Arboviruses Can Infect Wild Animals and Jump to Sylvatic Maintenance Cycles in South America.Front Cell Infect Microbiol. 2019 Jul 17;9:259. doi: 10.3389/fcimb.2019.00259. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31380302 Free PMC article. Review.
-
Aedes vittatus (Bigot) mosquito: An emerging threat to public health.J Vector Borne Dis. 2017 Oct-Dec;54(4):295-300. doi: 10.4103/0972-9062.225833. J Vector Borne Dis. 2017. PMID: 29460858 Review.
Cited by
-
Zika virus transmission by Brazilian Aedes aegypti and Aedes albopictus is virus dose and temperature-dependent.PLoS Negl Trop Dis. 2020 Sep 8;14(9):e0008527. doi: 10.1371/journal.pntd.0008527. eCollection 2020 Sep. PLoS Negl Trop Dis. 2020. PMID: 32898136 Free PMC article.
-
Potential Role of Accessory Domains in Polyproteins Encoded by Retrotransposons in Anti-viral Defense of Host Cells.Front Microbiol. 2019 Jan 4;9:3193. doi: 10.3389/fmicb.2018.03193. eCollection 2018. Front Microbiol. 2019. PMID: 30687243 Free PMC article. No abstract available.
-
Adventitious viruses persistently infect three commonly used mosquito cell lines.Virology. 2018 Aug;521:175-180. doi: 10.1016/j.virol.2018.06.007. Epub 2018 Jun 26. Virology. 2018. PMID: 29957338 Free PMC article.
-
Understanding the Mechanisms Underlying Host Restriction of Insect-Specific Viruses.Viruses. 2020 Aug 31;12(9):964. doi: 10.3390/v12090964. Viruses. 2020. PMID: 32878245 Free PMC article. Review.
-
Defective viral genomes as therapeutic interfering particles against flavivirus infection in mammalian and mosquito hosts.Nat Commun. 2021 Apr 16;12(1):2290. doi: 10.1038/s41467-021-22341-7. Nat Commun. 2021. PMID: 33863888 Free PMC article.
References
-
- World Health Organization (WHO). World Health Report. Executive Summary: Insect-borne diseases http://www.who.int/whr/1996/media_centre/executive_summary1/en/index9.html (1996).
-
- Adelman Z. N., Blair C. D., Carlson J. O., Beaty B. J. & Olson K. E. Sindbis virus-induced silencing of dengue viruses in mosquitoes. Insect Mol. Biol. 10, 265–273 (2001). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

