Lysine and the Na+/K+ Selectivity in Mammalian Voltage-Gated Sodium Channels
- PMID: 27584582
- PMCID: PMC5008630
- DOI: 10.1371/journal.pone.0162413
Lysine and the Na+/K+ Selectivity in Mammalian Voltage-Gated Sodium Channels
Abstract
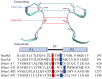
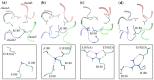
Voltage-gated sodium (Nav) channels are critical in the generation and transmission of neuronal signals in mammals. The crystal structures of several prokaryotic Nav channels determined in recent years inspire the mechanistic studies on their selection upon the permeable cations (especially between Na+ and K+ ions), a property that is proposed to be mainly determined by residues in the selectivity filter. However, the mechanism of cation selection in mammalian Nav channels lacks direct explanation at atomic level due to the difference in amino acid sequences between mammalian and prokaryotic Nav homologues, especially at the constriction site where the DEKA motif has been identified to determine the Na+/K+ selectivity in mammalian Nav channels but is completely absent in the prokaryotic counterparts. Among the DEKA residues, Lys is of the most importance since its mutation to Arg abolishes the Na+/K+ selectivity. In this work, we modeled the pore domain of mammalian Nav channels by mutating the four residues at the constriction site of a prokaryotic Nav channel (NavRh) to DEKA, and then mechanistically investigated the contribution of Lys in cation selection using molecular dynamics simulations. The DERA mutant was generated as a comparison to understand the loss of ion selectivity caused by the K-to-R mutation. Simulations and free energy calculations on the mutants indicate that Lys facilitates Na+/K+ selection by electrostatically repelling the cation to a highly Na+-selective location sandwiched by the carboxylate groups of Asp and Glu at the constriction site. In contrast, the electrostatic repulsion is substantially weakened when Lys is mutated to Arg, because of two intrinsic properties of the Arg side chain: the planar geometric design and the sparse charge distribution of the guanidine group.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Selective ion permeation involves complexation with carboxylates and lysine in a model human sodium channel.PLoS Comput Biol. 2018 Sep 12;14(9):e1006398. doi: 10.1371/journal.pcbi.1006398. eCollection 2018 Sep. PLoS Comput Biol. 2018. PMID: 30208027 Free PMC article.
-
The mechanism of Na⁺/K⁺ selectivity in mammalian voltage-gated sodium channels based on molecular dynamics simulation.Biophys J. 2013 Jun 4;104(11):2401-9. doi: 10.1016/j.bpj.2013.04.035. Biophys J. 2013. PMID: 23746512 Free PMC article.
-
Analysis of the selectivity filter of the voltage-gated sodium channel Na(v)Rh.Cell Res. 2013 Mar;23(3):409-22. doi: 10.1038/cr.2012.173. Epub 2012 Dec 18. Cell Res. 2013. PMID: 23247626 Free PMC article.
-
Simulation Studies of Ion Permeation and Selectivity in Voltage-Gated Sodium Channels.Curr Top Membr. 2016;78:215-60. doi: 10.1016/bs.ctm.2016.07.005. Epub 2016 Aug 3. Curr Top Membr. 2016. PMID: 27586286 Review.
-
The hitchhiker's guide to the voltage-gated sodium channel galaxy.J Gen Physiol. 2016 Jan;147(1):1-24. doi: 10.1085/jgp.201511492. J Gen Physiol. 2016. PMID: 26712848 Free PMC article. Review.
Cited by
-
Role of the Interaction Motif in Maintaining the Open Gate of an Open Sodium Channel.Biophys J. 2018 Nov 20;115(10):1920-1930. doi: 10.1016/j.bpj.2018.10.001. Epub 2018 Oct 4. Biophys J. 2018. PMID: 30366630 Free PMC article.
-
Purification and Characterization of JZTx-14, a Potent Antagonist of Mammalian and Prokaryotic Voltage-Gated Sodium Channels.Toxins (Basel). 2018 Oct 10;10(10):408. doi: 10.3390/toxins10100408. Toxins (Basel). 2018. PMID: 30308978 Free PMC article.
-
Selective ion permeation involves complexation with carboxylates and lysine in a model human sodium channel.PLoS Comput Biol. 2018 Sep 12;14(9):e1006398. doi: 10.1371/journal.pcbi.1006398. eCollection 2018 Sep. PLoS Comput Biol. 2018. PMID: 30208027 Free PMC article.
-
Molecular Dynamics Simulations of Ion Permeation in Human Voltage-Gated Sodium Channels.J Chem Theory Comput. 2023 May 23;19(10):2953-2972. doi: 10.1021/acs.jctc.2c00990. Epub 2023 Apr 28. J Chem Theory Comput. 2023. PMID: 37116214 Free PMC article.
-
Modeling the human Nav1.5 sodium channel: structural and mechanistic insights of ion permeation and drug blockade.Drug Des Devel Ther. 2017 Aug 4;11:2301-2324. doi: 10.2147/DDDT.S133944. eCollection 2017. Drug Des Devel Ther. 2017. PMID: 28831242 Free PMC article.
References
-
- Hille B. Ion channels of excitable membranes. Sunderland, MA: Sinauer Associates; 2001.
-
- Jiang Y, Lee A, Chen J, Ruta V, Cadene M, Chait BT, et al. X-ray structure of a voltage-dependent K+ channel. Nature. 2003;423(6935):33–41. . - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

