What have single-molecule studies taught us about gene expression?
- PMID: 27601529
- PMCID: PMC5024679
- DOI: 10.1101/gad.281725.116
What have single-molecule studies taught us about gene expression?
Abstract
The production of a single mRNA is the result of many sequential steps, from docking of transcription factors to polymerase initiation, elongation, splicing, and, finally, termination. Much of our knowledge about the fundamentals of RNA synthesis and processing come from ensemble in vitro biochemical measurements. Single-molecule approaches are very much in this same reductionist tradition but offer exquisite sensitivity in space and time along with the ability to observe heterogeneous behavior and actually manipulate macromolecules. These techniques can also be applied in vivo, allowing one to address questions in living cells that were previously restricted to reconstituted systems. In this review, we examine the unique insights that single-molecule techniques have yielded on the mechanisms of gene expression.
Keywords: fluorescence; single molecule; splicing; transcription.
Published by Cold Spring Harbor Laboratory Press.
Figures

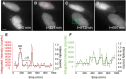


References
-
- Agafonov DE, Kastner B, Dybkov O, Hofele RV, Liu WT, Urlaub H, Luhrmann R, Stark H. 2016. Molecular architecture of the human U4/U6.U5 tri-snRNP. Science 351: 1416–1420. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
