The Potential Roles of the G1LEA and G3LEA Proteins in Early Embryo Development and in Response to Low Temperature and High Salinity in Artemia sinica
- PMID: 27603306
- PMCID: PMC5014412
- DOI: 10.1371/journal.pone.0162272
The Potential Roles of the G1LEA and G3LEA Proteins in Early Embryo Development and in Response to Low Temperature and High Salinity in Artemia sinica
Abstract
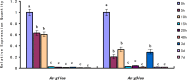
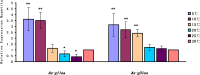
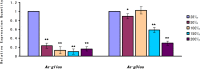
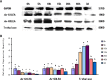
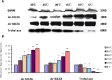
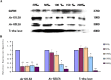
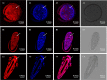
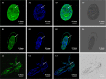
Late embryogenesis abundant proteins (LEA) are stress resistance-related proteins that play crucial roles in protecting against desiccation, cold and high salinity in a variety of animals and plants. However, the expression pattern, distribution and functions of LEA proteins in the post-diapause period of Artemia sinica, and under high salinity and low temperature stresses, remain unknown. In this study, the complete cDNA sequences of the group 1 LEA (As-g1lea) and group 3 LEA (As-g3lea) genes from A. sinica were cloned. The expression patterns and location of As-G1LEA and As-G1LEA were investigated. The protein abundances of As-G1LEA, As-G3LEA and Trehalase were analyzed during different developmental stages of the embryo and under low temperature and high salinity stresses in A. sinica. The full-length cDNA of As-g1lea was 960 bp, encoding a 182 amino acid protein, and As-g3lea was 2089 bp, encoding a 364 amino acid protein. As-g1lea and As-g3lea showed their highest expressions at 0 h of embryonic development and both showed higher relative expression in embryonic, rather than adult, development stages. The abundances of As-G1LEA, As-G3LEA and trehalose were upregulated under low temperature and downregulated under high salinity stress. These two genes did not show any tissue or organ specific expression. Our results suggested that these LEA proteins might play a pivotal role in stress tolerance in A. sinica.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
Cloning, expression pattern, and potential role of apoptosis inhibitor 5 in the termination of embryonic diapause and early embryo development of Artemia sinica.Gene. 2017 Sep 10;628:170-179. doi: 10.1016/j.gene.2017.07.021. Epub 2017 Jul 12. Gene. 2017. PMID: 28710039
-
Identification of the glycerol kinase gene and its role in diapause embryo restart and early embryo development of Artemia sinica.Gene. 2014 Mar 1;537(1):51-62. doi: 10.1016/j.gene.2013.12.029. Epub 2013 Dec 21. Gene. 2014. PMID: 24365596
-
Cloning and expression of retinoblastoma-binding protein 4 gene in embryo diapause termination and in response to salinity stress from brine shrimp Artemia sinica.Gene. 2016 Oct 15;591(2):351-61. doi: 10.1016/j.gene.2016.06.010. Epub 2016 Jun 5. Gene. 2016. PMID: 27267406
-
Stress tolerance during diapause and quiescence of the brine shrimp, Artemia.Cell Stress Chaperones. 2016 Jan;21(1):9-18. doi: 10.1007/s12192-015-0635-7. Epub 2015 Sep 3. Cell Stress Chaperones. 2016. PMID: 26334984 Free PMC article. Review.
-
Molecular chaperones, stress resistance and development in Artemia franciscana.Semin Cell Dev Biol. 2003 Oct;14(5):251-8. doi: 10.1016/j.semcdb.2003.09.019. Semin Cell Dev Biol. 2003. PMID: 14986854 Review.
Cited by
-
The Potential Roles of the Apoptosis-Related Protein PDRG1 in Diapause Embryo Restarting of Artemia sinica.Int J Mol Sci. 2018 Jan 2;19(1):126. doi: 10.3390/ijms19010126. Int J Mol Sci. 2018. PMID: 29301330 Free PMC article.
-
Transcriptomic analysis elucidates the molecular processes associated with hydrogen peroxide-induced diapause termination in Artemia-encysted embryos.PLoS One. 2021 Feb 19;16(2):e0247160. doi: 10.1371/journal.pone.0247160. eCollection 2021. PLoS One. 2021. PMID: 33606769 Free PMC article.
-
The Pyrus sinkiangensis Yu PsLEA4 Gene Enhances the Cold Resistance of Solanum lycopersicum.Plants (Basel). 2025 Jan 10;14(2):180. doi: 10.3390/plants14020180. Plants (Basel). 2025. PMID: 39861533 Free PMC article.
-
Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica.Genes (Basel). 2019 Sep 30;10(10):768. doi: 10.3390/genes10100768. Genes (Basel). 2019. PMID: 31574972 Free PMC article.
-
The Different Expression Patterns of HSP22, a Late Embryogenesis Abundant-like Protein, in Hypertrophic H9C2 Cells Induced by NaCl and Angiotensin II.Electrolyte Blood Press. 2018 Jun;16(1):1-10. doi: 10.5049/EBP.2018.16.1.1. Epub 2018 Jun 30. Electrolyte Blood Press. 2018. PMID: 30046328 Free PMC article.
References
-
- Kuenen DJ, Bass-Becking LGM. Historical notes on Artemia salina. Zoologischer Mededeelingen. 1938;20: 222–230.
-
- Anderson E, Lochhead JH, Lochhead MS, Huebner E. The origin and structure of the tertiary envelope in thick-shelled eggs of the brine shrimp, Artemia. J. Ultrastruct. Res. 1970;32, 497–525. - PubMed
-
- Clegg JS, Trotman CNA. Physiological and biochemical aspects of Artemia ecology. Artemia: Basic and Applied Biology. 2002;1:129–170.
-
- Jiang LJ, Hou L, Zou XY, Zhang RF, Wang J Q, Sun WJ, et al. Cloning and expression analysis of p26 gene in Artemia sinica. Acta Biochim Biophys Sinica. 200739 (5):351–358. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

