Single Nucleotide Polymorphism Discovery in Bovine Pituitary Gland Using RNA-Seq Technology
- PMID: 27606429
- PMCID: PMC5015895
- DOI: 10.1371/journal.pone.0161370
Single Nucleotide Polymorphism Discovery in Bovine Pituitary Gland Using RNA-Seq Technology
Abstract
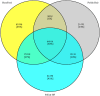
Examination of bovine pituitary gland transcriptome by strand-specific RNA-seq allows detection of putative single nucleotide polymorphisms (SNPs) within potential candidate genes (CGs) or QTLs regions as well as to understand the genomics variations that contribute to economic trait. Here we report a breed-specific model to successfully perform the detection of SNPs in the pituitary gland of young growing bulls representing Polish Holstein-Friesian (HF), Polish Red, and Hereford breeds at three developmental ages viz., six months, nine months, and twelve months. A total of 18 bovine pituitary gland polyA transcriptome libraries were prepared and sequenced using the Illumina NextSeq 500 platform. Sequenced FastQ databases of all 18 young bulls were submitted to NCBI-SRA database with NCBI-SRA accession numbers SRS1296732. For the investigated young bulls, a total of 113,882,3098 raw paired-end reads with a length of 156 bases were obtained, resulting in an approximately 63 million paired-end reads per library. Breed-wise, a total of 515.38, 215.39, and 408.04 million paired-end reads were obtained for Polish HF, Polish Red, and Hereford breeds, respectively. Burrows-Wheeler Aligner (BWA) read alignments showed 93.04%, 94.39%, and 83.46% of the mapped sequencing reads were properly paired to the Polish HF, Polish Red, and Hereford breeds, respectively. Constructed breed-specific SNP-db of three cattle breeds yielded at 13,775,885 SNPs. On an average 765,326 breed-specific SNPs per young bull were identified. Using two stringent filtering parameters, i.e., a minimum 10 SNP reads per base with an accuracy ≥ 90% and a minimum 10 SNP reads per base with an accuracy = 100%, SNP-db records were trimmed to construct a highly reliable SNP-db. This resulted in a reduction of 95,7% and 96,4% cut-off mark of constructed raw SNP-db. Finally, SNP discoveries using RNA-Seq data were validated by KASP™ SNP genotyping assay. The comprehensive QTLs/CGs analysis of 76 QTLs/CGs with RNA-seq data identified KCNIP4, CCSER1, DPP6, MAP3K5 and GHR CGs with highest SNPs hit loci in all three breeds and developmental ages. However, CAST CG with more than 100 SNPs hits were observed only in Polish HF and Hereford breeds.These findings are important for identification and construction of novel tissue specific SNP-db and breed specific SNP-db dataset by screening of putative SNPs according to QTL db and candidate genes for bovine growth and reproduction traits, one can develop genomic selection strategies for growth and reproductive traits.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
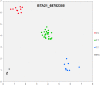
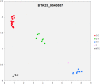

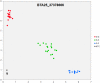
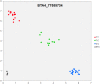

References
-
- Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. (2008). Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5: 1–8. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

