Atg5-dependent autophagy contributes to the development of acute myeloid leukemia in an MLL-AF9-driven mouse model
- PMID: 27607576
- PMCID: PMC5059867
- DOI: 10.1038/cddis.2016.264
Atg5-dependent autophagy contributes to the development of acute myeloid leukemia in an MLL-AF9-driven mouse model
Abstract
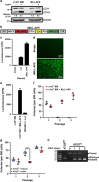
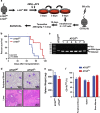
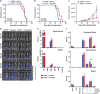
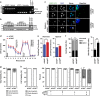
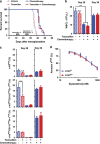
Acute myeloid leukemia (AML) is a hierarchical hematopoietic malignancy originating from leukemic stem cells (LSCs). Autophagy is a lysosomal degradation pathway that is hypothesized to be important for the maintenance of AML as well as contribute to chemotherapy response. Here we employ a mouse model of AML expressing the fusion oncogene MLL-AF9 and explore the effects of Atg5 deletion, a key autophagy protein, on the malignant transformation and progression of AML. Consistent with a transient decrease in colony-forming potential in vitro, the in vivo deletion of Atg5 in MLL-AF9-transduced bone marrow cells during primary transplantation prolonged the survival of recipient mice, suggesting that autophagy has a role in MLL-AF9-driven leukemia initiation. In contrast, deletion of Atg5 in malignant AML cells during secondary transplantation did not influence the survival or chemotherapeutic response of leukemic mice. Interestingly, autophagy was found to be involved in the survival of differentiated myeloid cells originating from MLL-AF9-driven LSCs. Taken together, our data suggest that Atg5-dependent autophagy may contribute to the development but not chemotherapy sensitivity of murine AML induced by MLL-AF9.
Figures
Similar articles
-
Differential requirement for wild-type Flt3 in leukemia initiation among mouse models of human leukemia.Exp Hematol. 2014 Mar;42(3):192-203.e1. doi: 10.1016/j.exphem.2013.11.008. Epub 2013 Nov 20. Exp Hematol. 2014. PMID: 24269847 Free PMC article.
-
EVI1 is critical for the pathogenesis of a subset of MLL-AF9-rearranged AMLs.Blood. 2012 Jun 14;119(24):5838-49. doi: 10.1182/blood-2011-11-393827. Epub 2012 May 2. Blood. 2012. PMID: 22553314 Free PMC article.
-
MLL-AF9 leukemia stem cells: hardwired or taking cues from the microenvironment?Cancer Cell. 2008 Jun;13(6):465-7. doi: 10.1016/j.ccr.2008.05.012. Cancer Cell. 2008. PMID: 18538728
-
Learning from mouse models of MLL fusion gene-driven acute leukemia.Biochim Biophys Acta Gene Regul Mech. 2020 Aug;1863(8):194550. doi: 10.1016/j.bbagrm.2020.194550. Epub 2020 Apr 19. Biochim Biophys Acta Gene Regul Mech. 2020. PMID: 32320749 Review.
-
Exploring the contribution of Zfp521/ZNF521 on primary hematopoietic stem/progenitor cells and leukemia progression.Cell Tissue Res. 2024 Dec;398(3):161-173. doi: 10.1007/s00441-024-03926-2. Epub 2024 Oct 22. Cell Tissue Res. 2024. PMID: 39436449 Free PMC article. Review.
Cited by
-
LncRNA LINC00265/miR-485-5p/IRF2-mediated autophagy suppresses apoptosis in acute myeloid leukemia cells.Am J Transl Res. 2020 Jun 15;12(6):2451-2462. eCollection 2020. Am J Transl Res. 2020. PMID: 32655783 Free PMC article.
-
Autophagy, cancer and angiogenesis: where is the link?Cell Biosci. 2019 Aug 13;9:65. doi: 10.1186/s13578-019-0327-6. eCollection 2019. Cell Biosci. 2019. PMID: 31428311 Free PMC article. Review.
-
Genetic polymorphisms of autophagy-related gene 5 (ATG5) rs473543 predict different disease-free survivals of triple-negative breast cancer patients receiving anthracycline- and/or taxane-based adjuvant chemotherapy.Chin J Cancer. 2018 Jan 31;37(1):4. doi: 10.1186/s40880-018-0268-1. Chin J Cancer. 2018. PMID: 29382381 Free PMC article.
-
Advances in Understanding the Links between Metabolism and Autophagy in Acute Myeloid Leukemia: From Biology to Therapeutic Targeting.Cells. 2023 Jun 5;12(11):1553. doi: 10.3390/cells12111553. Cells. 2023. PMID: 37296673 Free PMC article. Review.
-
Current Understanding of the Role of Autophagy in the Treatment of Myeloid Leukemia.Int J Mol Sci. 2024 Nov 14;25(22):12219. doi: 10.3390/ijms252212219. Int J Mol Sci. 2024. PMID: 39596291 Free PMC article. Review.
References
-
- Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med 2015; 373: 1136–1152. - PubMed
-
- Somervaille TCP, Cleary ML. Identification and characterization of leukemia stem cells in murine MLL-AF9 acute myeloid leukemia. Cancer Cell 2006; 10: 257–268. - PubMed
-
- Barabé F, Kennedy JA, Hope KJ, Dick JE. Modeling the initiation and progression of human acute leukemia in mice. Science 2007; 316: 600–604. - PubMed
-
- Corral J, Lavenir I, Impey H, Warren AJ, Forster A, Larson TA et al. An Mll-AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: a method to create fusion oncogenes. Cell 1996; 85: 853–861. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

